 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Seita.4G270600.1.p | ||||||||
| Common Name | LOC101786720, SETIT_006354mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Cenchrinae; Setaria
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 474aa MW: 51795.4 Da PI: 5.7608 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.4 | 5.9e-32 | 267 | 321 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
k rlrWt eLHerFveav++L G+ekAtPk +l+lmkv+gLt++hvkSHLQkYRl
Seita.4G270600.1.p 267 KSRLRWTLELHERFVEAVNKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRL 321
68****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 11.767 | 264 | 324 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 8.24E-17 | 265 | 321 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.1E-30 | 265 | 322 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.3E-24 | 267 | 322 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.0E-8 | 269 | 320 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 3.9E-22 | 357 | 403 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 474 aa Download sequence Send to blast |
MSTQSVATGE QIIAPNETVH ACTSTQTSVL QLFDSKSDHR LLIDDTLSST SQSSSIKTEL 60 IRSSSLSRSL SVNLQKRSPE TDPESPLSHI SHPKFSDPIL SNSSTFCTSL FSSSSKNTDP 120 CRQMGTLPFL PHPPKCEQQV SAGQSSSSSL LFAGDTGNAL DEAEHSDDLK DFLNLSGDAS 180 DGSFHGETNA LAFDEQMEFQ FLSEQLGIAI TDNEESPHLD DIYGTPPQLS SLPVSSCSNQ 240 SIQNLGSPVK VQLSSSRSSS VSATTNKSRL RWTLELHERF VEAVNKLEGP EKATPKGVLK 300 LMKVEGLTIY HVKSHLQKYR LAKYLPETKE DEKASSEDKK AQSGSSSSDS SKTKNLQVAE 360 ALRMQMEVQK QLHEQLEVQR QLQLRIEEHA RYLQKILEEQ QKAGNLSLKA PTKAQAVSPE 420 STASKERSET EAGTSSPRPS KNRNLDAHSE CKSPAVSKRT EFQVDPESEV PCS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 2e-25 | 267 | 324 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 2e-25 | 267 | 324 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 2e-25 | 267 | 324 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 2e-25 | 267 | 324 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 2e-25 | 267 | 324 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 2e-25 | 267 | 324 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 2e-25 | 267 | 324 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 2e-25 | 267 | 324 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in phosphate starvation signaling (PubMed:26082401). Binds to P1BS, an imperfect palindromic sequence 5'-GNATATNC-3', to promote the expression of inorganic phosphate (Pi) starvation-responsive genes (PubMed:26082401). Functionally redundant with PHR1 and PHR2 in regulating Pi starvation response and Pi homeostasis (PubMed:26082401). {ECO:0000269|PubMed:26082401}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00354 | DAP | Transfer from AT3G13040 | Download |
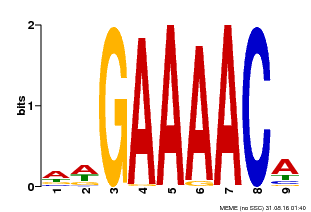
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Seita.4G270600.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated under Pi starvation conditions. {ECO:0000269|PubMed:26082401}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004966348.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 isoform X1 | ||||
| Swissprot | Q6YXZ4 | 1e-165 | PHR3_ORYSJ; Protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| TrEMBL | K3XWP6 | 0.0 | K3XWP6_SETIT; Uncharacterized protein | ||||
| STRING | Si006354m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4526 | 37 | 69 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G13040.2 | 2e-67 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Seita.4G270600.1.p |
| Entrez Gene | 101786720 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



