 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SapurV1A.0985s0080.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Salix
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 339aa MW: 36882.5 Da PI: 9.2803 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 38.5 | 2.6e-12 | 83 | 127 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT+ E++++++a +++ + Wk+I + +g +t q++s+ qky
SapurV1A.0985s0080.2.p 83 RESWTEPEHDKFLEALQLFDRD-WKKIGAFIG-SKTIIQIRSHAQKY 127
789*****************77.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.2E-15 | 77 | 133 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 14.574 | 78 | 132 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 2.1E-17 | 81 | 130 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 3.0E-6 | 81 | 133 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.1E-9 | 82 | 130 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 5.5E-10 | 83 | 127 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.57E-7 | 85 | 128 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 339 aa Download sequence Send to blast |
MQNPLIQTLK TPQQKPISST SMVSKNPNPA DGSYLDPTGM AALPGLSPFA TTTTPLTTTT 60 TSSSTSEDPS KKIRKPYTIT KSRESWTEPE HDKFLEALQL FDRDWKKIGA FIGSKTIIQI 120 RSHAQKYFLK VQKSGTNEHL PPPRPKRKAT HPYPQKASKN AIVPSQPSDA FQSSSAPLEP 180 GYVLRPDSSS IPVDPIPSAA VASSWTNNVP TVSLSNQTKG PVAANNCCNG TESTPRTKLT 240 GKTAEQGNHG HSMRVLPDFS QVYGFIGSVF DPNVTDQLQN LKKMDPIDVE TVLLLMRNLS 300 LNLNSPSFEE HRMLLSTHET DSESIGANNN VDQSMNVA* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00625 | PBM | Transfer from PK02532.1 | Download |
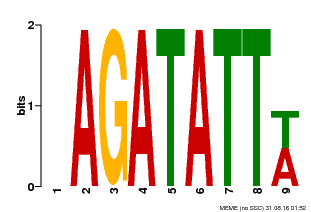
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | SapurV1A.0985s0080.2.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002305938.2 | 0.0 | protein REVEILLE 6 | ||||
| Swissprot | Q8H0W3 | 1e-125 | RVE6_ARATH; Protein REVEILLE 6 | ||||
| TrEMBL | A0A3N7EYR6 | 0.0 | A0A3N7EYR6_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0004s07160.1 | 0.0 | (Populus trichocarpa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G52660.1 | 1e-110 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | SapurV1A.0985s0080.2.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



