 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SapurV1A.0849s0010.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Salix
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 324aa MW: 35763.3 Da PI: 8.1162 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 55.1 | 1.7e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd +lv +++++G+g+W++++ g+ R+ k+c++rw +yl
SapurV1A.0849s0010.1.p 14 KGPWTPEEDIILVSYIQEHGPGNWRAVPTNTGLLRCSKSCRLRWTNYL 61
79******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 47.2 | 5e-15 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T E++ ++++ ++lG++ W++Ia++++ Rt++++k++w+++l
SapurV1A.0849s0010.1.p 67 RGNFTDHEEKMIIHLQALLGNR-WAAIASYLP-QRTDNDIKNFWNTHL 112
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.9E-23 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 24.347 | 9 | 65 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 6.36E-30 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.0E-14 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.3E-16 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.09E-11 | 16 | 61 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 3.2E-24 | 65 | 116 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 8.5E-15 | 66 | 114 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 18.674 | 66 | 116 | IPR017930 | Myb domain |
| Pfam | PF00249 | 1.7E-13 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 8.39E-11 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0010468 | Biological Process | regulation of gene expression | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 324 aa Download sequence Send to blast |
MGRPPCCDKI GVKKGPWTPE EDIILVSYIQ EHGPGNWRAV PTNTGLLRCS KSCRLRWTNY 60 LRPGIKRGNF TDHEEKMIIH LQALLGNRWA AIASYLPQRT DNDIKNFWNT HLKKKLRKIQ 120 AGQGDQSRDG LSSAGSQQIS RGQWERRLQT DINMARQALC EALSPGKPSS LLTGLKPACV 180 NEKPAPAPIY ASSTENIARL LKGWMINGPK QSLKNSTTTQ NSFINMAGAD LLSGEGTPDK 240 ADKNGTGLSQ TFESLFGFDS FDSSNSDFSQ PMSPDTGLFQ DESKPNSGGQ VPLSLIERWL 300 FDEGAMQGKD YLNEVTIDED NLF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1mse_C | 6e-25 | 14 | 116 | 4 | 105 | C-Myb DNA-Binding Domain |
| 1msf_C | 6e-25 | 14 | 116 | 4 | 105 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. {ECO:0000305}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00579 | DAP | Transfer from AT5G62470 | Download |
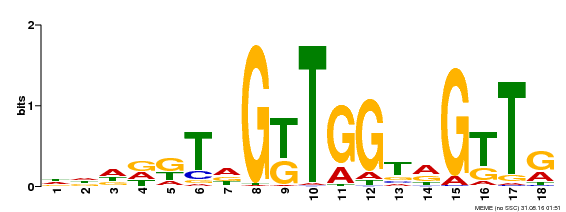
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | SapurV1A.0849s0010.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002323853.1 | 0.0 | myb-related protein 306 | ||||
| Swissprot | P81392 | 1e-117 | MYB06_ANTMA; Myb-related protein 306 | ||||
| TrEMBL | B9IJX2 | 0.0 | B9IJX2_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0017s11880.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1016 | 34 | 115 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62470.2 | 1e-110 | myb domain protein 96 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | SapurV1A.0849s0010.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



