 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SapurV1A.0805s0030.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Salix
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 839aa MW: 92231.5 Da PI: 6.3802 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.7 | 2.1e-18 | 18 | 76 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++ +++ps +r++L +++ +++ +q+kvWFqNrR +ek+
SapurV1A.0805s0030.1.p 18 KYVRYTPEQVEALERLYHDCPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 76
5679*****************************************************97 PP
| |||||||
| 2 | START | 173.4 | 1.4e-54 | 162 | 370 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg 88
+aee+++e+++ka+ ++ Wv+++ +++g+++ +++ s++++g +ra+g+v +++ v+e+l+d++ W ++++++++l+v+ ++
SapurV1A.0805s0030.1.p 162 IAEETLTEFLSKATGTAVEWVQMPGMKPGPDSNGIVAISHGCTGVGARACGLVGLEPT-RVAEILKDRPSWFRDCRAVDVLNVLPTA 247
789*******************************************************.8888888888****************99 PP
..EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE- CS
START 89 ..galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwveh 169
g+++l +++l+a+++l+p Rdf+ +Ry+ l++g++v++++S+ ++q+ p+ +++vRae+lpSg+l++p+++g+s +++v+h
SapurV1A.0805s0030.1.p 248 ngGTIELLYMQLYAPTTLAPgRDFWLLRYTSVLEDGSLVVCERSLKNTQNGPSmppVQHFVRAEMLPSGYLVRPCEGGGSIIHIVDH 334
99*************************************************9988899***************************** PP
EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 170 vdlkgrlphwllrslvksglaegaktwvatlqrqce 205
+dl+ ++++++lr+l++s+++ ++kt++a+l+++++
SapurV1A.0805s0030.1.p 335 MDLEPWSVPEVLRPLYESSTVLAQKTTMAALRQLRQ 370
********************************9876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.613 | 13 | 77 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.2E-15 | 15 | 81 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.37E-16 | 17 | 81 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.31E-16 | 18 | 78 | No hit | No description |
| Pfam | PF00046 | 5.8E-16 | 19 | 76 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 3.5E-18 | 20 | 76 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 1.23E-6 | 70 | 109 | No hit | No description |
| PROSITE profile | PS50848 | 25.535 | 152 | 367 | IPR002913 | START domain |
| CDD | cd08875 | 5.77E-77 | 156 | 372 | No hit | No description |
| SMART | SM00234 | 8.1E-38 | 161 | 371 | IPR002913 | START domain |
| Gene3D | G3DSA:3.30.530.20 | 2.1E-22 | 161 | 368 | IPR023393 | START-like domain |
| SuperFamily | SSF55961 | 5.36E-36 | 161 | 373 | No hit | No description |
| Pfam | PF01852 | 4.4E-52 | 162 | 370 | IPR002913 | START domain |
| Pfam | PF08670 | 2.0E-52 | 695 | 837 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 839 aa Download sequence Send to blast |
MAMSCKDGKN LINMDNGKYV RYTPEQVEAL ERLYHDCPKP SSIRRQQLIR ECPILSNIEP 60 KQIKVWFQNR RCREKQRKEA SRLQAVNRKL TAMNKLLMEE NDRLQKQVSQ LVYENGYFRQ 120 HTQNTTLASK DTSCESVVTS GQHHLTPQHP PRDASPAGLL SIAEETLTEF LSKATGTAVE 180 WVQMPGMKPG PDSNGIVAIS HGCTGVGARA CGLVGLEPTR VAEILKDRPS WFRDCRAVDV 240 LNVLPTANGG TIELLYMQLY APTTLAPGRD FWLLRYTSVL EDGSLVVCER SLKNTQNGPS 300 MPPVQHFVRA EMLPSGYLVR PCEGGGSIIH IVDHMDLEPW SVPEVLRPLY ESSTVLAQKT 360 TMAALRQLRQ IAQEASQSNV TNWGRRPAAL RALSQRLSRG FNEALNGFSD EGWSMIGNDG 420 MDDVTILVNS SPDKLMGLNL PFSNGFPAVS SAVLCAKASM LLQNVPPAIL LRFLREHRSE 480 WADNNIDAYA AAAVKVGPCS LQGSRVGNFG GQVILPLAHT IEQEEFLEVI KLEGVCHSPE 540 DAIMPRDVFL LQLCCGMDEN AVGTCAELIF APIDATFADE APLLPSGFRI IPLDSGKEAS 600 SPNRTLDLAS ALEVGAGNRA SSGFSANSGC TRSVMTIAFE FAFESNMQEH VASMARQYIR 660 SIISSVQRVA LALSPSHLGS QAGLRSPLGT PEAQTLAHWI CQSYRGYLGV ELLKSSSEGS 720 ESILKTLWHH SDAIMCCSLK ALPVFTFANQ AGLDMLETTL VALQDITLEK IFDDHGRKTL 780 CSEFPQIMQQ GFTCLQGGIC MSSMGRPVSY ERAVSWKVLN EEENAHCICF MFINWSFV* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
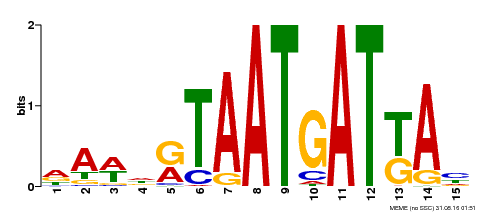
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | SapurV1A.0805s0030.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KF534716 | 0.0 | KF534716.1 Populus alba x Populus glandulosa HD-ZIPIII mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024464085.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A2I4KKW3 | 0.0 | A0A2I4KKW3_POPTR; HD-ZIP family protein | ||||
| STRING | POPTR_0001s18930.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1946 | 33 | 87 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | SapurV1A.0805s0030.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



