 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SapurV1A.0767s0100.4.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Salix
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 730aa MW: 84021.1 Da PI: 8.2227 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 86.1 | 5.5e-27 | 87 | 189 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk............tekerrtraetrtgCkaklkvkkekdgk 75
+fY+eYA+++GF++ +++s++sk+++e+++++f Cs++g+++e +k+ e+ +r+ ++t+Cka+++vk++ dgk
SapurV1A.0767s0100.4.p 87 SFYQEYARSMGFNTAIQNSRRSKTSREFIDAKFACSRYGTKREYDKSfnrprsrqtkqdPENGTGRRSCSKTDCKASMHVKRRPDGK 173
5*******************************************999999998776555444555999******************* PP
FAR1 76 wevtkleleHnHelap 91
w+++++++eHnHel p
SapurV1A.0767s0100.4.p 174 WVIHSFVKEHNHELLP 189
*************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 7.8E-25 | 87 | 189 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 1.6E-31 | 287 | 378 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.322 | 567 | 603 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.4E-7 | 578 | 605 | IPR006564 | Zinc finger, PMZ-type |
| Pfam | PF04434 | 1.0E-4 | 579 | 602 | IPR007527 | Zinc finger, SWIM-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 730 aa Download sequence Send to blast |
MDINLRLPSG DHDKEGEEPN GVNNMLSEAK LHNGDVEIGN VVDVAEQVLS IEGGDVNSPT 60 TSMGFKEDIK LEPLSGMEFE SHGAAYSFYQ EYARSMGFNT AIQNSRRSKT SREFIDAKFA 120 CSRYGTKREY DKSFNRPRSR QTKQDPENGT GRRSCSKTDC KASMHVKRRP DGKWVIHSFV 180 KEHNHELLPA QAVSEQTRKM YAAMAQQFAE YKNVVGLKND PKNPFDKGWN LGLEAGETKI 240 LLDFFTRMQN MNSNFFYAVD LGEDQRLKNL IWADAKSRHD YSNFSDVVNF DTTYVRNKYK 300 MPLALFVGVN QHYQFMLLGC ALVSDESAAT YSWLMQTWLR AMGGQTPKVI ITDQDKVLKQ 360 VISEVFPNAH HCFCLWNVLG KVSENLGNVI KQNRNFMAKF DKCIFRSWTE NEFGKRWLKI 420 LDRFELRENE WMQSLYEDRE QWVPIYMRGA FLAGMSTALR SESINSYFDK YVHKKTTVQE 480 FVRQYESILQ DRYEEEAKAD SDTWNKQPAL KSPSPLEKSV SGMYTHAVFK KFQVEVLGVV 540 ACHPKMESQD ETSVSFRVQD LEKEQDFTVL WNQMRLEVSC ICRLYEYKGY LCRHALVVLQ 600 MCQQSAIPSQ YILKRWTKDA KSRHLVGEEC EQVQSRVQRY NDLCQRALKL SEEASLSQES 660 YSMAFRALEE AFGNCIGMNN SNKNLVEAGT SATHGLLCIE DDNQNRSVTK TIKKKNQTKK 720 RKVGGQKQI* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
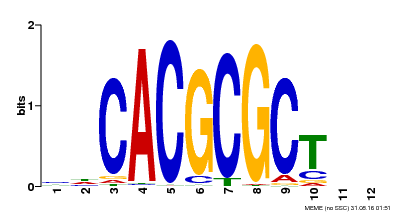
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | SapurV1A.0767s0100.4.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024459461.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X2 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A2K1ZVK4 | 0.0 | A0A2K1ZVK4_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0006s02150.1 | 0.0 | (Populus trichocarpa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | SapurV1A.0767s0100.4.p |



