 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SapurV1A.0767s0090.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Salix
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 839aa MW: 96649.3 Da PI: 7.1301 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 90.1 | 2.9e-28 | 73 | 160 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnH 87
+fY+eYAk++GF++++++s++sk+++e+++++f Cs++g + e+++ ++r++++++t+Cka+++vk++ dgkw ++++++eHnH
SapurV1A.0767s0090.2.p 73 SFYQEYAKSMGFTTSIKNSRRSKKSKEFIDAKFACSRYGVTPESDSG---NSRRSTVKKTDCKASMHVKRRPDGKWIIHEFVKEHNH 156
5****************************************999888...888999******************************* PP
FAR1 88 elap 91
el p
SapurV1A.0767s0090.2.p 157 ELLP 160
*975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 1.1E-25 | 73 | 160 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 2.4E-24 | 281 | 373 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.248 | 560 | 596 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 2.4E-4 | 570 | 594 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 9.2E-7 | 571 | 598 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 839 aa Download sequence Send to blast |
MTERNHKIID DMIDLQDNVP ADDVVGGNIV DVVHSRDVAV VDSPKRAVVM FEGDVNYELC 60 DGIEFGSHEE AYSFYQEYAK SMGFTTSIKN SRRSKKSKEF IDAKFACSRY GVTPESDSGN 120 SRRSTVKKTD CKASMHVKRR PDGKWIIHEF VKEHNHELLP ALAYHFRIHR NVKLAEKNNI 180 DILHAVSERT RKMYVEMSRQ SGGYQNFGFV KSELNMQFEK GRHLALDEGD AQVVLEYFKH 240 VKKENANFFY AIDLNEDQRL RNLFWVDSKS REDYISFNDA VCFETFYVKY HEKLPFAPFV 300 GVNHHSQPIL LGCAFIADES RSSFVWLMKT WLRAMGGQAP KVIVTDVDKT LKAAIEEAFP 360 NTRHCFSLWH ILERLPETLS HVIKRHEKFL PKFHKCIFKS WTDDRFDMRW WKMVTQFELQ 420 DDEWIQSLYE DRKKWVPTYL GDTFLAGTSA AQRSESMSAF FDKYIHRKIT MKEFMKQYGT 480 ILQNRYEDES VADFDTSHKQ PALKSPSPWE KQMSLVYTHA IFKKFQVEVL GVVGCHPKRE 540 SEDGTLVTFR VQDCEKDEQF LVTWNQTKSE VSCFCHLFEY KGFLCRHALI VLQICGLSTI 600 PPHYILKRWT KDAKSRQPLA VGTERAQTRA QRYNDLCKLA IEMSEEGSLS EESYNIVLHT 660 LVEALKNCVN VNNCNNSVAE SSTFTLTHRE AEEENQGSLV TKSSKKKNPV RKRKVQSDPD 720 VMLVEAPENL QQMENLSSEG INLSGYYGTQ QNVQGLVQLN LMEPPHDGYY VNQQSMQGLG 780 QLNSIAPSHD GFFGNQQSMH GLGQFDFRPP SGFSYSMQLQ DDTHLRSSHM HGGASRNA* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
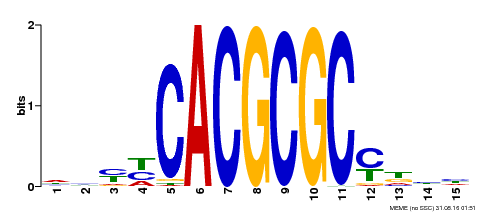
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | SapurV1A.0767s0090.2.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024459852.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 isoform X1 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A2K1ZVH1 | 0.0 | A0A2K1ZVH1_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0006s02140.1 | 0.0 | (Populus trichocarpa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | SapurV1A.0767s0090.2.p |



