 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SapurV1A.0561s0300.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Salix
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 505aa MW: 55645.8 Da PI: 6.6775 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 158.5 | 2.8e-49 | 19 | 145 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp.kkvkaeekewyfFskrdkkyatgkrknratksgyWkat 86
+ pGfrF Ptd el+++yLkkk+eg + + evi+e++i+k+ePwdLp k+v +e+ew+fFs r +ky++g++++rat++gyWkat
SapurV1A.0561s0300.1.p 19 MFPGFRFTPTDVELISYYLKKKIEGAERCV-EVISEIEICKHEPWDLPaKSVMPSESEWFFFSARGRKYPNGSQSKRATEHGYWKAT 104
579************************777.99***************666667899****************************** PP
NAM 87 gkdkevlskkgelvglkktLvfykgrapkgektdWvmheyrl 128
gk+++v+s ++e++g+k+tLvf+ grapkge+t+W+mhey +
SapurV1A.0561s0300.1.p 105 GKERNVKS-GSEVIGTKRTLVFHAGRAPKGERTEWIMHEYCM 145
********.9******************************87 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 4.45E-54 | 10 | 160 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 52.329 | 19 | 161 | IPR003441 | NAC domain |
| Pfam | PF02365 | 6.7E-25 | 21 | 144 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009845 | Biological Process | seed germination | ||||
| GO:0009938 | Biological Process | negative regulation of gibberellic acid mediated signaling pathway | ||||
| GO:0033619 | Biological Process | membrane protein proteolysis | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0071472 | Biological Process | cellular response to salt stress | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005886 | Cellular Component | plasma membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 505 aa Download sequence Send to blast |
MATGGVSPET QMSIEASSMF PGFRFTPTDV ELISYYLKKK IEGAERCVEV ISEIEICKHE 60 PWDLPAKSVM PSESEWFFFS ARGRKYPNGS QSKRATEHGY WKATGKERNV KSGSEVIGTK 120 RTLVFHAGRA PKGERTEWIM HEYCMQGQSQ DSLVVCRLRK NAEFRPSDTS NRGSVNETHL 180 STTHSSGDAV SDGGIGQGGV PSREKAAECS KSNDSCSIEQ LSSASESEQK LSNDAALAET 240 SSHQKDSDDE EDCFADILKH DIIKLDEVLL SSPPGLRSKV ASNPEAKTRS EQPGEIFISE 300 ALPLPSIPLQ GTSKQPVKIF MSASLPLHST PSQGTEHQPV EMFISEALPL HSTPSRGTEH 360 RPVEMFMSEA LPLHSTPSRG TEHQPVEMFM SETLPLHSTP SQGTEHQPVE VFMSEALPLP 420 STPSQGTASR RIMLRRQMPG MSQAEMHTNN VDRESKLPVH CAKKGKLRHI SAVFILLALL 480 VSLIAGFKQV ERIIYASMYR TSWH* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 3e-41 | 1 | 160 | 4 | 164 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 3e-41 | 1 | 160 | 4 | 164 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 3e-41 | 1 | 160 | 4 | 164 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 3e-41 | 1 | 160 | 4 | 164 | NO APICAL MERISTEM PROTEIN |
| 4dul_A | 3e-41 | 1 | 160 | 4 | 164 | NAC domain-containing protein 19 |
| 4dul_B | 3e-41 | 1 | 160 | 4 | 164 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator activated by proteolytic cleavage through regulated intramembrane proteolysis (RIP), probably via metalloprotease activity. Regulates gibberellic acid-mediated salt-responsive repression of seed germination and flowering via FT, thus delaying seed germination under high salinity conditions. {ECO:0000269|PubMed:17410378, ECO:0000269|PubMed:18363782, ECO:0000269|PubMed:19704528, ECO:0000269|PubMed:19704545}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
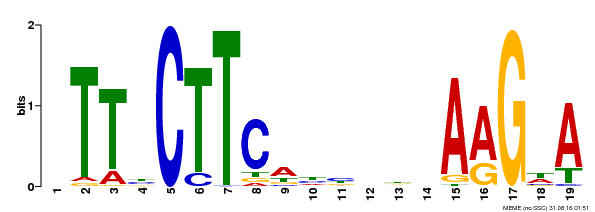
| Motif ID | Method | Source | Motif file |
| MP00280 | DAP | Transfer from AT2G27300 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | SapurV1A.0561s0300.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By high salt stress (PubMed:19704545, PubMed:17410378, PubMed:18363782, PubMed:19704528). Repressed by gibberellic acid (GA), but induced by the GA biosynthetic inhibitor paclabutrazol (PAC) (PubMed:18363782). Accumulates transiently in seeds upon imbibition (PubMed:19704545, PubMed:17410378, PubMed:18363782, PubMed:19704528). Induced by drought stress (PubMed:17158162). {ECO:0000269|PubMed:17158162, ECO:0000269|PubMed:17410378, ECO:0000269|PubMed:18363782, ECO:0000269|PubMed:19704528, ECO:0000269|PubMed:19704545}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011043268.1 | 0.0 | PREDICTED: NAC domain-containing protein 89-like isoform X1 | ||||
| Swissprot | Q9XIN7 | 4e-96 | NAC40_ARATH; NAC domain-containing protein 40 | ||||
| TrEMBL | A0A2K1Z8S2 | 0.0 | A0A2K1Z8S2_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0009s16290.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7315 | 33 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G27300.1 | 2e-98 | NTM1-like 8 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | SapurV1A.0561s0300.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



