 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SMil_00025905-RA_Salv | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lamiaceae; Nepetoideae; Mentheae; Salvia
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 347aa MW: 39029.7 Da PI: 9.7462 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 41.2 | 4.1e-13 | 46 | 94 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s++kGV + +grW A+I++ ++r++lg+f ++eAa+a++ a+++++g
SMil_00025905-RA_Salv 46 SRFKGVVPQP-NGRWGAQIYEK-----HQRVWLGTFNEEDEAARAYDTAAQRFRG 94
689***9788.8*********3.....5**********99*************98 PP
| |||||||
| 2 | B3 | 104.5 | 5.6e-33 | 166 | 267 | 1 | 98 |
EEEE-..-HHHHTT-EE--HHH.HTT....---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh....ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkl 84
f+k+ tpsdv+k++rlv+pk++ae+h +g++ +++ l +ed +g++W+++++y+++s++yvltkGW++Fvk+++L++gD+v F++
SMil_00025905-RA_Salv 166 FEKAVTPSDVGKLNRLVIPKQHAEKHfplqSGNSSKGVLLNFEDAKGKVWRFRYSYWNSSQSYVLTKGWSRFVKEKDLRAGDIVSFQR 253
89****************************888899***************************************************8 PP
-SSSEE..EEEEE- CS
B3 85 dgrsefelvvkvfr 98
+ +++l++ + +
SMil_00025905-RA_Salv 254 STGPDKQLYIDWKE 267
86688888887765 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 2.6E-8 | 46 | 94 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.87E-23 | 46 | 101 | No hit | No description |
| SuperFamily | SSF54171 | 9.15E-17 | 46 | 102 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 4.5E-20 | 47 | 102 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.3E-29 | 47 | 108 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.875 | 47 | 102 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 5.8E-43 | 158 | 276 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 8.24E-30 | 164 | 264 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.24E-28 | 165 | 254 | No hit | No description |
| SMART | SM01019 | 1.2E-24 | 166 | 269 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.7E-29 | 166 | 268 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 14.635 | 166 | 269 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 347 aa Download sequence Send to blast |
MDGSSVDDST TTDTLSMTLA SPSPSPPAKS PDLDGVEAES RKLPSSRFKG VVPQPNGRWG 60 AQIYEKHQRV WLGTFNEEDE AARAYDTAAQ RFRGRDAVTN FKPLSDDEAE AAFLSSHSKA 120 EIVDMLRKHT YADELEQSRR SYGAGGSRTA ERDGERAAAK AREQLFEKAV TPSDVGKLNR 180 LVIPKQHAEK HFPLQSGNSS KGVLLNFEDA KGKVWRFRYS YWNSSQSYVL TKGWSRFVKE 240 KDLRAGDIVS FQRSTGPDKQ LYIDWKERNN NGTASNPVQV MRLFGVNIFQ ADYCSGKRMR 300 EMEIMGLDCT KKQRIINVLG GGEGGVRWRC RNLQNLQLYS KIDKKEG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 4e-59 | 163 | 269 | 11 | 118 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional repressor of flowering time on long day plants. Acts directly on FT expression by binding 5'-CAACA-3' and 5'-CACCTG-3 sequences. Functionally redundant with TEM2. {ECO:0000269|PubMed:18718758}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00024 | PBM | Transfer from AT1G25560 | Download |
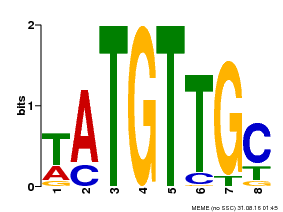
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Expressed with a circadian rhythm showing a peak at dawn. {ECO:0000269|PubMed:18718758}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012846076.1 | 1e-177 | PREDICTED: LOW QUALITY PROTEIN: AP2/ERF and B3 domain-containing transcription factor RAV1-like | ||||
| Swissprot | Q9C6M5 | 1e-143 | RAVL1_ARATH; AP2/ERF and B3 domain-containing transcription repressor TEM1 | ||||
| TrEMBL | A0A4D9A8D6 | 0.0 | A0A4D9A8D6_SALSN; RAV-like factor | ||||
| STRING | Migut.E00306.1.p | 1e-172 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2958 | 24 | 53 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25560.1 | 1e-142 | RAV family protein | ||||



