 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SMil_00017002-RA_Salv | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lamiaceae; Nepetoideae; Mentheae; Salvia
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 343aa MW: 39030.6 Da PI: 5.3689 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 100.6 | 1e-31 | 63 | 116 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtpeLH rFv+ave+LGG+++AtPk +l+lm++kgL ++hvkSHLQ+YR+
SMil_00017002-RA_Salv 63 PRLRWTPELHLRFVHAVERLGGQDRATPKLVLQLMGIKGLNIAHVKSHLQMYRS 116
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 13.211 | 59 | 119 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.7E-30 | 59 | 117 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 3.4E-15 | 61 | 118 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.5E-22 | 63 | 118 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.8E-9 | 64 | 115 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 343 aa Download sequence Send to blast |
MEISEKECLK RSPCEGMDDE DESEEQNELS FEEAKDGGSS SNSTVEESDK KPPVRPYVRS 60 KMPRLRWTPE LHLRFVHAVE RLGGQDRATP KLVLQLMGIK GLNIAHVKSH LQMYRSKKTD 120 ESNQGLKFME GVDRNIFNLS QLPLLPSFNQ RPNSNLRYGD PTWMHNSAIG LNSRSITRPG 180 FYAERFLSSH YTRLPSLNGS AWRNDDTRLK EEKLGLIHKQ EPWLGQTIQN PIEIDSLKQK 240 LQEKRSSPPN LTQILSQERE KLGLKRKASD AELLDLNLSL GLESRNDDED DDDDDDDGDE 300 HLGLSLYTFP EPSKDRVAKK MKGDFVNVGD EKARGASTLD LTL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 1e-17 | 64 | 118 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 1e-17 | 64 | 118 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 1e-17 | 64 | 118 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 1e-17 | 64 | 118 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
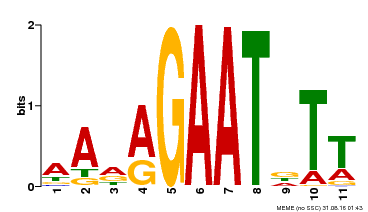
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00297 | DAP | Transfer from AT2G38300 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011095220.1 | 1e-125 | uncharacterized protein LOC105174731 | ||||
| TrEMBL | A0A4D9AME3 | 1e-157 | A0A4D9AME3_SALSN; Uncharacterized protein | ||||
| STRING | Migut.J00123.1.p | 1e-100 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4382 | 21 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38300.1 | 2e-53 | G2-like family protein | ||||



