 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Rsa1.0_02220.1_g00001.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 566aa MW: 62280.2 Da PI: 4.8036 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.5 | 9.9e-13 | 395 | 440 | 4 | 54 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksL 54
+h e+Er+RR+++N++f Lr ++P+ K++Ka+ L A+ YI++L
Rsa1.0_02220.1_g00001.1 395 NHVEAERQRREKLNQRFYSLRAVVPNV-----SKMDKASLLGDAISYINEL 440
799***********************6.....5***************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 1.3E-54 | 64 | 252 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 17.282 | 391 | 440 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 3.93E-18 | 394 | 458 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 3.72E-15 | 394 | 445 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 1.4E-18 | 395 | 459 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.6E-10 | 395 | 440 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.2E-16 | 397 | 446 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006952 | Biological Process | defense response | ||||
| GO:0009718 | Biological Process | anthocyanin-containing compound biosynthetic process | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043425 | Molecular Function | bHLH transcription factor binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 566 aa Download sequence Send to blast |
MDDHRSRVSL SPPDPHMSNY FLNQTTPPAD NDNAVEAFIG TNHWSQQTQP PPPQTLSQFN 60 EDTLQQRLQT LIESAGERWT YAIFWQISHE FDSPTGENTV ILGWGDGYYR GEEDKEKKKK 120 QSLSSNPAEQ EHRKRVIREL NSLIAGGGGA GVSDEANDEE VTDTEWFFLV SMTQSFVNGV 180 GLPGESFLNS RVIWLSGSGA LTGSGCERAR QGEVYGLQTI VCIAAENGVV ELGSSEVISQ 240 SSDLMDKVNG LFNGNGNGEA SSWGFNLNPD QGENDPALWL SEPNVTGIEP VVQETPPPGA 300 IENAGGDLNF SSSGLNQSGN FKQGSSSSKK RSPVVSKDEE MLSFTTVVRS AAKSVDSDHS 360 DIEASVVKEA IIVEPEKKPR KRGRKPANGR EEPLNHVEAE RQRREKLNQR FYSLRAVVPN 420 VSKMDKASLL GDAISYINEL KTKLQQAEAD KEEVQKQLDG MGKEGGGSRR AKERKSNRDS 480 ESSVEMEIDV KIIGWDVMIR VQCGKKNHPG ARFMEALKEL DLEVNHASLS VVNDLMIQQA 540 TVKMGSQFFN HDQLKAALML KVGEDI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4rru_A | 1e-109 | 34 | 248 | 17 | 227 | Transcription factor MYC3 |
| 4ywc_A | 1e-109 | 34 | 248 | 17 | 227 | Transcription factor MYC3 |
| 4ywc_B | 1e-109 | 34 | 248 | 17 | 227 | Transcription factor MYC3 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 376 | 384 | KKPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in tryptophan, jasmonic acid (JA) and other stress-responsive gene regulation. With MYC2 and MYC4, controls additively subsets of JA-dependent responses. Can form complexes with all known glucosinolate-related MYBs to regulate glucosinolate biosynthesis. Binds to the G-box (5'-CACGTG-3') of promoters. Activates multiple TIFY/JAZ promoters. {ECO:0000269|PubMed:12136026, ECO:0000269|PubMed:21242320, ECO:0000269|PubMed:21321051, ECO:0000269|PubMed:21335373, ECO:0000269|PubMed:23943862}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00085 | PBM | Transfer from AT5G46760 | Download |
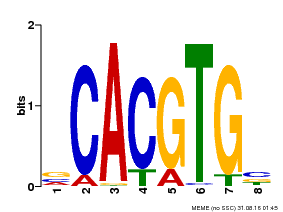
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Rsa1.0_02220.1_g00001.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Barely up-regulated by jasmonic acid. {ECO:0000269|PubMed:21242320, ECO:0000269|PubMed:21335373}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018457039.1 | 0.0 | PREDICTED: transcription factor MYC3 | ||||
| Swissprot | Q9FIP9 | 0.0 | MYC3_ARATH; Transcription factor MYC3 | ||||
| TrEMBL | A0A397XV80 | 0.0 | A0A397XV80_BRACM; Uncharacterized protein | ||||
| TrEMBL | A0A3P6E5A0 | 0.0 | A0A3P6E5A0_BRAOL; Uncharacterized protein | ||||
| STRING | Bo9g065530.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1265 | 28 | 96 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G46760.1 | 0.0 | bHLH family protein | ||||



