|
Plant Transcription
Factor Database
|
Transcription Factor Information
|
Basic
Information? help
Back to Top |
| TF ID |
Rsa1.0_01567.1_g00002.1 |
| Organism |
|
| Taxonomic ID |
|
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
| Family |
HD-ZIP |
| Protein Properties |
Length: 689aa MW: 76296.5 Da PI: 6.9581 |
| Description |
HD-ZIP family protein |
| Gene Model |
| Gene Model ID |
Type |
Source |
Coding Sequence |
| Rsa1.0_01567.1_g00002.1 | genome | RGD | View CDS |
|
| Signature Domain? help Back to Top |
 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | Homeobox | 59.2 | 6.5e-19 | 45 | 98 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k +++t++q++eLe++F+ +++p++++r eL +kl+L+ +q+k+WFqNrR+++k
Rsa1.0_01567.1_g00002.1 45 KYRRHTSYQIQELESFFKVCPHPNEKQRLELGSKLSLESKQIKFWFQNRRTQMK 98
56789**********************************************999 PP
|
| 2 | START | 168.2 | 5.5e-53 | 210 | 428 | 3 | 205 |
HHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS...SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....E CS
START 3 aeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv..dsgealrasgvvdmvlallveellddkeqWdetla....k 78
a a++el+++a+++ p+W k s +s + de+++ f+ sk + e +r++g+v ++ +lve+l++++ +W e+++ +
Rsa1.0_01567.1_g00002.1 210 AVVAMDELIRLAEVDNPLWIKCSkserDSMKHDEYTSLFAGSKHpgFAAEGSRETGLVLINSLTLVETLMNTN-RWAEMFEcivaA 294
56689******************88888888999999998888778888889*********************.************ PP
EEEEEEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEE CS
START 79 aetlevissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepk 157
a+t+evis+g g lqlm ae+q++splvp R + f+Ry++q+g+g w++vdvS d++++ + +s+ ++lpSg++ie+
Rsa1.0_01567.1_g00002.1 295 ASTVEVISNGsggsrnGSLQLMEAEFQVMSPLVPiRQVKFLRYCKQHGDGLWAVVDVSYDVNRESQDLKSYDGLKRLPSGCIIEDI 380
****************************************************************99899999999*********** PP
CTCEEEEEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 158 snghskvtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqce 205
+ng skvtw+eh +++g+++h l+++l++s++ ga +w+atlqr ce
Rsa1.0_01567.1_g00002.1 381 GNGCSKVTWIEHSEYEGSHIHPLYQQLLSSSVGLGATKWLATLQRRCE 428
***********************************************9 PP
|
| Sequence ? help Back to Top |
| Protein Sequence Length: 689 aa
Download sequence Send
to blast |
MNGDLDITKG DDEFESDSLE EMSGDDDDNK HEQRPKKKKK KKMSKYRRHT SYQIQELESF 60
FKVCPHPNEK QRLELGSKLS LESKQIKFWF QNRRTQMKTQ LERHENAILR QENEKLRVEN 120
GILKEAMRSP TCTNCGGAAT PGEVSHEQQQ LRMENAKLKY ELDKLCALAN RFIGGSISLE 180
QPSNGGGVGS QDLSLGHGFT RGTSTFMDLA VVAMDELIRL AEVDNPLWIK CSKSERDSMK 240
HDEYTSLFAG SKHPGFAAEG SRETGLVLIN SLTLVETLMN TNRWAEMFEC IVAAASTVEV 300
ISNGSGGSRN GSLQLMEAEF QVMSPLVPIR QVKFLRYCKQ HGDGLWAVVD VSYDVNRESQ 360
DLKSYDGLKR LPSGCIIEDI GNGCSKVTWI EHSEYEGSHI HPLYQQLLSS SVGLGATKWL 420
ATLQRRCESY TTLLSSQDQT GLSLAGTKST QTLAQRMKLS FYSGITASPI HKWEKLVAEN 480
VGQDTRILTR RSLQPSGVVL SAATSMWLPV TQQRLFEFLC DSKCRNQWDI LCNGASVENM 540
LLIPKRQREG RCVSLLQPAG KHQSGNSMLI LQETWSDASG ALVVYAPVDV PSMNMVMSGG 600
DSANVALLPS GFSISPDGSS SSSSSWYDQI DTHGGFVNHE SKGCLLTVGF QILVNSVPTT 660
KLNMESVQTV NNLIACTIHK IKAALSIPA
|
| Functional Description ? help
Back to Top |
| Source |
Description |
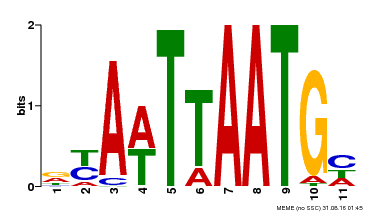
| UniProt | Probable transcription factor that binds to the DNA sequence 5'-GCATTAAATGC-3'. {ECO:0000269|PubMed:16778018}. |