 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Rsa1.0_00685.1_g00005.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 830aa MW: 95393.7 Da PI: 6.7879 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 52 | 2.4e-16 | 86 | 173 | 2 | 91 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnH 87
fY+eY+++vGF++ +++s++sk+++e+++++f Cs++g+++e +k+ +++ r r++++ + + + dgkw++++++ +HnH
Rsa1.0_00685.1_g00005.1 86 FYQEYSRTVGFNTAIQNSRRSKTTREFIDAKFACSRYGTKREYDKSFNNRPRARQSKQHQE--HPPPPXKPDGKWVIHSFVRDHNH 169
******************************************9999555555555555554..444555668************** PP
FAR1 88 elap 91
el p
Rsa1.0_00685.1_g00005.1 170 ELLP 173
*975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 1.2E-13 | 86 | 173 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 3.3E-26 | 273 | 364 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.161 | 553 | 589 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 2.9E-6 | 564 | 591 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 830 aa Download sequence Send to blast |
MDIDLRLHSG DLCKEAGDDD EQQQQQHRGL DNEDIDISKI EDDVTVDVQH SDNNNNNNTS 60 VPEQQGVNLE PLNGMEFNSH GEAYTFYQEY SRTVGFNTAI QNSRRSKTTR EFIDAKFACS 120 RYGTKREYDK SFNNRPRARQ SKQHQEHPPP PXKPDGKWVI HSFVRDHNHE LLPAQAVSEQ 180 TRKVYAAMAK QFAEYKTVIS LKSESKGSFE KGRSLSVVET GDFKVLLEFL SRMQVLNSNF 240 FYAVDLGDDD QRVKSVFWVD AKSRFNYGSF CDVVSLDTTY VRNKYKMPLA VFVGVNQHYQ 300 YMVLGCALLS DESGATFSWL METWLRAMGG QAPKVLITEQ DAVLNSVVPE VFPSTQHRLF 360 LWHVLIKVSE NLGQVAKQHE NFTAKFEKCI YKSWKEEDFA RRWWKILARF GLKDDQWMQS 420 LYEDRRKWAP TYMTDVLLAG MSTSQRADSV NAYFDKYLHK KTSVQEFVKL YDMILQDRFE 480 EEAKADSETW NKPPTMKSPS PFEKSVSEVY TPAVFKKFQI EVLGAIACCP REENRDATCS 540 TFRVQDFENN QDFVVTWNQA KAEVSCMCRL FEYKGYLCRH TLNVLQCCHL SSIPSQYILK 600 RWTKDARSQH FPGEPQQLQT RVQRYNDLCQ RALKLSEEAS ISQESYNITF LAIEEALGNC 660 AGVNTSGRSL PDVVTSPTQG LISVEEDNQS RSAVKTTSKK KNPTKKRKVN SEQEVMPVAA 720 PESLQPMDKL SPRTVGLESY YGTQQSVQNL VQLNLMGPNR DNFYGNHQQT IQGLRQLNSI 780 APSYDSYYTA QQGIHGQGVD FFRPPNFTYD IRDDPNVRTP QLHEDASRHT |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
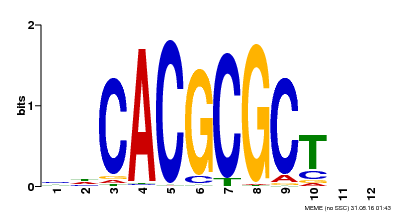
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Rsa1.0_00685.1_g00005.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018492466.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X1 | ||||
| Refseq | XP_018492467.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X1 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A3P5YVM0 | 0.0 | A0A3P5YVM0_BRACM; Uncharacterized protein | ||||
| TrEMBL | M4ERA7 | 0.0 | M4ERA7_BRARP; Uncharacterized protein | ||||
| STRING | Bra031330.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM118 | 26 | 253 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||



