 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Rsa1.0_00324.1_g00001.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 342aa MW: 37931.5 Da PI: 9.8707 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 56.1 | 9.2e-18 | 118 | 167 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
s+y+GV++++++grW+++I+d + k+++lg f+ta+ Aa+a+++a+ k++g
Rsa1.0_00324.1_g00001.1 118 SQYRGVTFYRRTGRWESHIWD------CgKQVYLGGFDTAYAAARAYDRAAIKFRG 167
78*******************......55************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 2.35E-15 | 118 | 175 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 1.9E-9 | 118 | 167 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 17.332 | 119 | 175 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.73E-11 | 119 | 174 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 1.4E-16 | 119 | 175 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 4.9E-30 | 119 | 181 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009658 | Biological Process | chloroplast organization | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 342 aa Download sequence Send to blast |
MLDLSLGMFS NYGEDQDRKV PLVSAAGEEE SNISSSSSSS KDSAAGKAFI TFGIIKRDDD 60 VVPPPPSKET GDLFPVADAR RNIEFSLDDG HWLRLSSLQR NKQQAVKKSR RGPRSRSSQY 120 RGVTFYRRTG RWESHIWDCG KQVYLGGFDT AYAAARAYDR AAIKFRGLDA DINFVVEDYK 180 QDLDKMKNLN KVEFVQTLRR ESASFGRGSS KYKGLTLQKC TQFKTHQDQI HLLQNRGWDA 240 AAIKYSELGK GGVMKSGVHI KGNAHNDLEL SLGLSSSSQN AKLTTRDYYE GINQSTMGLF 300 SKQSSIYLPV TTMTPLKTVA ASSGFPFITM TTSSSSTTSF DP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). Repressor of flowering. {ECO:0000250, ECO:0000269|PubMed:14573523}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00075 | ChIP-chip | Transfer from AT3G54990 | Download |
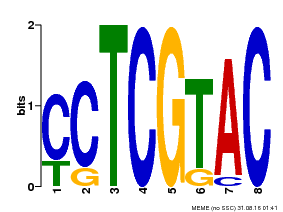
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Rsa1.0_00324.1_g00001.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Repressed by miR172 after photoperiod changement. {ECO:0000269|PubMed:14573523}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018491037.1 | 0.0 | PREDICTED: AP2-like ethylene-responsive transcription factor SMZ | ||||
| Swissprot | Q6PV68 | 1e-158 | SMZ_ARATH; AP2-like ethylene-responsive transcription factor SMZ | ||||
| TrEMBL | A0A078JMS1 | 0.0 | A0A078JMS1_BRANA; BnaA09g54740D protein | ||||
| TrEMBL | A0A397Y5Y2 | 0.0 | A0A397Y5Y2_BRACM; Uncharacterized protein | ||||
| TrEMBL | M4CSD1 | 0.0 | M4CSD1_BRARP; Uncharacterized protein | ||||
| STRING | Bra007123.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM10858 | 18 | 32 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G54990.1 | 1e-153 | AP2 family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



