 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | RrC6965_p3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | MIKC_MADS | ||||||||
| Protein Properties | Length: 279aa MW: 32155.7 Da PI: 9.7564 | ||||||||
| Description | MIKC_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 99.6 | 1.2e-31 | 9 | 58 | 1 | 50 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeys 50
krienk+nrqvtfskRrng+lKKA+ELSvLCdaeva+i+fss+gklye+
RrC6965_p3 9 KRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIVFSSRGKLYEFG 58
79**********************************************95 PP
| |||||||
| 2 | K-box | 97.5 | 2e-32 | 104 | 198 | 6 | 99 |
K-box 6 gks.leeakaeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaLrkkle 99
+++ e+++a +++qe++kLk+++e+L r++RhllGedL ++s+keLq Le++Le++l+ R++K+++++e++e+l+kke++l ++nk+L+ k+e
RrC6965_p3 104 NNRpEESTQACNWSQEVTKLKAKYESLVRTNRHLLGEDLRDMSVKELQALERKLETALTATRQRKTQVMMEEMEDLRKKERQLGDINKQLKIKFE 198
33367888999********************************************************************************9987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50066 | 33.389 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| SMART | SM00432 | 6.3E-42 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 1.31E-29 | 2 | 64 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 6.39E-39 | 2 | 59 | No hit | No description |
| PRINTS | PR00404 | 1.4E-32 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 2.5E-27 | 10 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.4E-32 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.4E-32 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 3.1E-30 | 108 | 197 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 15.137 | 113 | 203 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009553 | Biological Process | embryo sac development | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010094 | Biological Process | specification of carpel identity | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0010582 | Biological Process | floral meristem determinacy | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048455 | Biological Process | stamen formation | ||||
| GO:0048459 | Biological Process | floral whorl structural organization | ||||
| GO:0048509 | Biological Process | regulation of meristem development | ||||
| GO:0048833 | Biological Process | specification of floral organ number | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0080112 | Biological Process | seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 279 aa Download sequence Send to blast |
MGRGRVEMKR IENKINRQVT FSKRRNGLLK KAYELSVLCD AEVALIVFSS RGKLYEFGSV 60 GYIVIRESDQ AFILWVFLKV KSFILRVGRT IERYHRCYNS SLTNNRPEES TQACNWSQEV 120 TKLKAKYESL VRTNRHLLGE DLRDMSVKEL QALERKLETA LTATRQRKTQ VMMEEMEDLR 180 KKERQLGDIN KQLKIKFEAE GLAFKSFQDL WPNSAASVAD DPNNSGFPVQ PSHPSSLDCN 240 TEPFLQIGFQ QHYYVQGEGS SVSKSNVARE TNFVQGWVL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5f28_A | 9e-19 | 1 | 60 | 1 | 60 | MEF2C |
| 5f28_B | 9e-19 | 1 | 60 | 1 | 60 | MEF2C |
| 5f28_C | 9e-19 | 1 | 60 | 1 | 60 | MEF2C |
| 5f28_D | 9e-19 | 1 | 60 | 1 | 60 | MEF2C |
| 6byy_A | 1e-18 | 1 | 60 | 1 | 60 | MEF2 CHIMERA |
| 6byy_B | 1e-18 | 1 | 60 | 1 | 60 | MEF2 CHIMERA |
| 6byy_C | 1e-18 | 1 | 60 | 1 | 60 | MEF2 CHIMERA |
| 6byy_D | 1e-18 | 1 | 60 | 1 | 60 | MEF2 CHIMERA |
| 6bz1_A | 1e-18 | 1 | 60 | 1 | 60 | MEF2 CHIMERA |
| 6bz1_B | 1e-18 | 1 | 60 | 1 | 60 | MEF2 CHIMERA |
| 6bz1_C | 1e-18 | 1 | 60 | 1 | 60 | MEF2 CHIMERA |
| 6bz1_D | 1e-18 | 1 | 60 | 1 | 60 | MEF2 CHIMERA |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. Forms a heterodimer via the K-box domain with AG, that could be involved in genes regulation during floral meristem development. | |||||
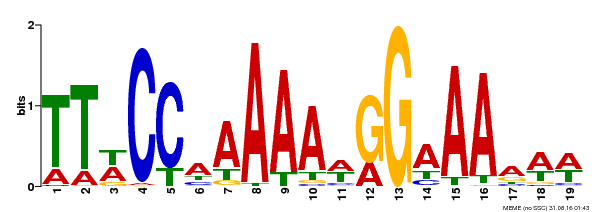
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00315 | DAP | Transfer from AT2G45650 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AJ508409 | 0.0 | AJ508409.1 Brassica oleracea var. botrytis mRNA for MADS-box protein AGL6-a (agl6-a gene), short splice variant. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018437530.1 | 0.0 | PREDICTED: agamous-like MADS-box protein AGL6 | ||||
| Swissprot | P29386 | 1e-160 | AGL6_ARATH; Agamous-like MADS-box protein AGL6 | ||||
| TrEMBL | Q8GTE8 | 1e-159 | Q8GTE8_BRAOB; MADS-box protein AGL6-a | ||||
| STRING | Bo4g024840.1 | 1e-159 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4302 | 26 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G45650.1 | 1e-163 | AGAMOUS-like 6 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



