 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | RrC450_p2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 749aa MW: 82323.6 Da PI: 6.1227 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 65.3 | 8.4e-21 | 127 | 182 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe++F+++++p++++r +L+k+l L++rqVk+WFqNrR+++k
RrC450_p2 127 KKRYHRHTPQQIQELESMFKECPHPDEKQRLDLSKRLCLETRQVKFWFQNRRTQMK 182
688999***********************************************999 PP
| |||||||
| 2 | START | 189.7 | 1.5e-59 | 311 | 519 | 1 | 194 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS...SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEEEECTT.. CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv..dsgealrasgvvdmvlallveellddkeqWdetla....kaetlevissg.. 88
ela++a++elvk+a+++ep+Wvks +++n de++++f+++k +ea+r+sg+v+ ++ lve+l+d++ +W+e+++ +a+t++ is g
RrC450_p2 311 ELALAAMDELVKLAQSDEPLWVKSLdgerDELNHDEYMRTFSSNKQtgLVSEASRTSGMVIINSLALVETLMDSN-RWTEMFPcnvaRATTTDIISGGma 409
5899*************************************99998999**************************.************************ PP
....EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--..-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHH CS
START 89 ....galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe.sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllr 182
galqlm+aelq+lsplvp R++ f+R+++q+ +g+w++vdvS+d ++++ s+++vR +lpSg++++++sng+skvtwveh++++++++h l+r
RrC450_p2 410 gtrsGALQLMNAELQVLSPLVPvRNVNFLRFCKQHAEGVWAVVDVSIDTVRDNSGvSAGMVR--RLPSGCVVQDMSNGYSKVTWVEHAEYDENQIHHLYR 507
******************************************************99999999..************************************ PP
HHHHHHHHHHHH CS
START 183 slvksglaegak 194
+l++s + g +
RrC450_p2 508 PLLRSAITPGGR 519
***998765544 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.64E-20 | 112 | 184 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 6.3E-22 | 115 | 184 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.038 | 124 | 184 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.6E-17 | 125 | 188 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 5.31E-18 | 126 | 184 | No hit | No description |
| Pfam | PF00046 | 2.5E-18 | 127 | 182 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 159 | 182 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 36.488 | 302 | 514 | IPR002913 | START domain |
| CDD | cd08875 | 7.80E-100 | 308 | 529 | No hit | No description |
| SuperFamily | SSF55961 | 1.74E-25 | 308 | 513 | No hit | No description |
| SMART | SM00234 | 1.9E-32 | 311 | 531 | IPR002913 | START domain |
| Pfam | PF01852 | 3.5E-52 | 311 | 515 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.44E-23 | 516 | 735 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 749 aa Download sequence Send to blast |
MNFGSLFDNT TNGGSPTGAR ILSGLTYGNA STVVPGGAMA QVAASSLFTP PINTKSVYSS 60 SGLSLALEQP ERGGTSMRNN NNDNFEGSTT NNRRSREEQE HESRSGSDNV GGISGEDQDA 120 DDKPPRKKRY HRHTPQQIQE LESMFKECPH PDEKQRLDLS KRLCLETRQV KFWFQNRRTQ 180 MKTQLERHEN ALLRQENDKL RAENMSIREA MRNPICNNCG GPAMLGDVSL EEHHLRIENA 240 RLKDELDRVC NLTGKFLGHH HNHSSSLELA VGNNNGGNFS FPPDFGLPPP QQQQHQPTVV 300 NGIDQRTVLL ELALAAMDEL VKLAQSDEPL WVKSLDGERD ELNHDEYMRT FSSNKQTGLV 360 SEASRTSGMV IINSLALVET LMDSNRWTEM FPCNVARATT TDIISGGMAG TRSGALQLMN 420 AELQVLSPLV PVRNVNFLRF CKQHAEGVWA VVDVSIDTVR DNSGVSAGMV RRLPSGCVVQ 480 DMSNGYSKVT WVEHAEYDEN QIHHLYRPLL RSAITPGGRK SMLKLAQRMT VNFCSGISAP 540 SVHSWSKLTV ENVDPDVRVM TRKSVDDSGI VLSAATSVWL PASPQRLFDF LRNERMRCEW 600 DILSNGGPMQ EMAHIAKGQD QGNSVSLLRS NPMNTNQSSM LILQETCIDA SGALVVYAPV 660 DIPAMNVVMN GGDSSYVALL PSGFAVLPDG GSGDVEQRPV GGGSLLTVAF QILVNNLPTA 720 KLTVESVETV NNLISCTVQK IRDALQCES |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
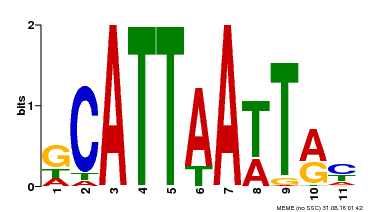
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK353253 | 0.0 | AK353253.1 Thellungiella halophila mRNA, complete cds, clone: RTFL01-28-B24. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018475317.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A3P6AA33 | 0.0 | A0A3P6AA33_BRACM; Uncharacterized protein | ||||
| STRING | Bo3g054630.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1128 | 27 | 105 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



