 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | RrC2237_p4 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 230aa MW: 26339.6 Da PI: 7.6581 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.6 | 2.1e-18 | 78 | 130 | 4 | 56 |
-SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 4 RttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+ ++t+ ql Le+ F+++ +++ +++ +L+++lgL+ rq+ vWFqNrRa++k
RrC2237_p4 78 KKRLTSGQLASLERSFQEEIKLDSDRKLKLSRELGLQPRQIAVWFQNRRARWK 130
45699***********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 115.2 | 3.8e-37 | 77 | 167 | 2 | 92 |
HD-ZIP_I/II 2 kkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelk 92
kk+rl++ q+++LE+sF+ee kL+++rK +l+reLglqprq+avWFqnrRAR+k+kqlE+ y++L+++yd +++e+++L++ev++Lr+ l+
RrC2237_p4 77 KKKRLTSGQLASLERSFQEEIKLDSDRKLKLSRELGLQPRQIAVWFQNRRARWKAKQLEQLYDSLRQEYDVVSREKQMLHEEVRKLRALLR 167
9**************************************************************************************8775 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.99E-18 | 64 | 134 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 16.649 | 72 | 132 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 3.0E-16 | 74 | 136 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.03E-14 | 77 | 133 | No hit | No description |
| Pfam | PF00046 | 5.4E-16 | 78 | 130 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 6.4E-19 | 79 | 139 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 5.6E-5 | 103 | 112 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 107 | 130 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 5.6E-5 | 112 | 128 | IPR000047 | Helix-turn-helix motif |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010434 | Biological Process | bract formation | ||||
| GO:0010582 | Biological Process | floral meristem determinacy | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 230 aa Download sequence Send to blast |
MEWTTTRNVE DVRIAFMTPP WPESSSFNSV HSSSYDPYAG SSYAPADTQL GPVTSVPESE 60 KIISAYQFSS NNNEMIKKKR LTSGQLASLE RSFQEEIKLD SDRKLKLSRE LGLQPRQIAV 120 WFQNRRARWK AKQLEQLYDS LRQEYDVVSR EKQMLHEEVR KLRALLRDHG LIKKQIPGGE 180 NTTEIPSVVI AHPGAENLNN NQISGENQIY GVDQYNNPML IVSSCWPPFP |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 124 | 132 | RRARWKAKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Putative transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
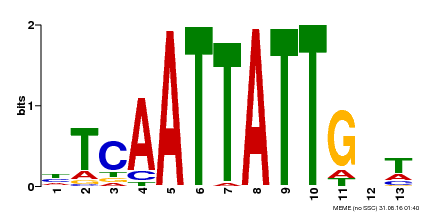
| Motif ID | Method | Source | Motif file |
| MP00486 | DAP | Transfer from AT5G03790 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KF939590 | 1e-107 | KF939590.1 Cardamine hirsuta LMI1 (LMI1) gene, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018444845.1 | 1e-171 | PREDICTED: putative homeobox-leucine zipper protein ATHB-51 | ||||
| Swissprot | Q9LZR0 | 1e-137 | ATB51_ARATH; Putative homeobox-leucine zipper protein ATHB-51 | ||||
| TrEMBL | A0A0D3EI30 | 1e-143 | A0A0D3EI30_BRAOL; Uncharacterized protein | ||||
| STRING | Bo9g181720.1 | 1e-143 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1660 | 28 | 88 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G03790.1 | 1e-131 | homeobox 51 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



