 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | RrC126_p9 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 853aa MW: 95127.7 Da PI: 6.9374 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 70.2 | 2.7e-22 | 162 | 263 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvv 94
f+k+lt sd++++g +++ +++a+e+ + + + +++l+ +d+++ +W++++i+r++++r++l++GW+ Fv++++L +gD ++F + +++el+v
RrC126_p9 162 FCKTLTASDTSTHGGFSVLRRHADEClppldmSRQ-PPTQELVAKDLHSIEWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL--RGENGELRV 258
99*****************************7333.34459************************************************..449999*** PP
EEE-S CS
B3 95 kvfrk 99
+v+r+
RrC126_p9 259 GVRRA 263
**996 PP
| |||||||
| 2 | Auxin_resp | 120.8 | 1e-39 | 288 | 370 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha+st+++F+v+Y+Pr+s+seF+v+++++++++k+++s+GmRfkm+fe+e+++e+r++Gt+vg++d+dp+rW +SkWrsLk
RrC126_p9 288 AWHAISTGTMFTVYYKPRTSPSEFIVPFDQYMESVKNNYSIGMRFKMRFEGEEAPEQRFTGTIVGIEDSDPTRWAKSKWRSLK 370
89*******************************************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 4.0E-40 | 158 | 277 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 3.66E-40 | 158 | 291 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.87E-20 | 160 | 262 | No hit | No description |
| PROSITE profile | PS50863 | 11.237 | 162 | 264 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 2.4E-22 | 162 | 264 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 4.1E-20 | 162 | 263 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 7.7E-37 | 288 | 370 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 4.8E-5 | 717 | 763 | IPR033389 | AUX/IAA domain |
| PROSITE profile | PS51745 | 24.774 | 727 | 811 | IPR000270 | PB1 domain |
| SuperFamily | SSF54277 | 1.14E-11 | 728 | 804 | No hit | No description |
| Pfam | PF02309 | 6.9E-6 | 773 | 812 | IPR033389 | AUX/IAA domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 853 aa Download sequence Send to blast |
MTSSEVSMKG NRGGGGGENF SSLGYSDSTT VAGEAMKTQS NRSVAAERIV DAEAALYREL 60 WHACAGPLVT VPRQDDRVFY FPQGHIEQVE ASTNQAAEQQ MPLYDLPSKI LCRVINVDLK 120 AEADTDEVYA QITLLPEPIQ DENAIEKVPP PPPPPRFQVH SFCKTLTASD TSTHGGFSVL 180 RRHADECLPP LDMSRQPPTQ ELVAKDLHSI EWRFRHIFRG QPRRHLLQSG WSVFVSSKRL 240 VAGDAFIFLR GENGELRVGV RRAMRQQGNV PSSVISSHSM HLGVLATAWH AISTGTMFTV 300 YYKPRTSPSE FIVPFDQYME SVKNNYSIGM RFKMRFEGEE APEQRFTGTI VGIEDSDPTR 360 WAKSKWRSLK VRWDETTSIP RPDRVSPWKI EPALSPPALS PVPMPRPKRP RSNIAPSTPD 420 SSMRIREGSS KANMDPLPAS GLSRVLQGQE YPTLRTKHVE SVECDAPENS IVWQSSNDDD 480 KVDVVSASRR YENWMSSGRH EQPAYTDLLS GFGANIEPPP HSGHQIPLYD HRLSSSPSVA 540 ARKILSDQDG KFEYLANQWQ MMHSGLSLKL HESPKVPAAS DTSFQGISNP YALPRAVTTE 600 NATGNWPIRP RALNYFEEAV HAQAREQHVT KRPAVVQEEA AKPRDGNCRL FGIPLVNNVN 660 GTDTAMAQRN NLNDTTGLTQ IASPKVQDLS DQSKGSKSTN DHREQGRPFP VSKPHPKDVQ 720 TKTNSSRSCT KVHKQGIALG RSVDLSKFQN YEELVTELDR LFEFNGELMA PKKDWLIVYT 780 DDENDMMLVG DDPWQEFCCM VRKIFIYTKE EVRKMNPGTL SYRNEEDAIA GEGSDAKDAK 840 SASNPSLSSA GNS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-178 | 55 | 392 | 18 | 354 | Auxin response factor 1 |
| 4ldw_A | 1e-178 | 55 | 392 | 18 | 354 | Auxin response factor 1 |
| 4ldw_B | 1e-178 | 55 | 392 | 18 | 354 | Auxin response factor 1 |
| 4ldx_A | 1e-178 | 55 | 392 | 18 | 354 | Auxin response factor 1 |
| 4ldx_B | 1e-178 | 55 | 392 | 18 | 354 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Could act as transcriptional activator or repressor. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Promotes flowering, stamen development, floral organ abscission and fruit dehiscence. Functions independently of ethylene and cytokinin response pathways. May act as a repressor of cell division and organ growth. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:15960614, ECO:0000269|PubMed:16176952, ECO:0000269|PubMed:16339187}. | |||||
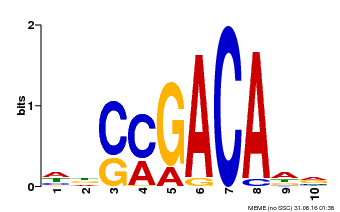
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JN979460 | 0.0 | JN979460.1 Brassica rapa subsp. pekinensis auxin response factor 2-2 (ARF2-2) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018491253.1 | 0.0 | PREDICTED: auxin response factor 2-like | ||||
| Swissprot | Q94JM3 | 0.0 | ARFB_ARATH; Auxin response factor 2 | ||||
| TrEMBL | H9B4C1 | 0.0 | H9B4C1_BRARP; Auxin response factor | ||||
| STRING | Bra010048.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5348 | 26 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||



