 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pp3c8_2690V3.3.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Bryophytina; Bryopsida; Funariidae; Funariales; Funariaceae; Physcomitrella
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1141aa MW: 127134 Da PI: 6.8811 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 171.7 | 1.1e-53 | 23 | 139 | 4 | 117 |
CG-1 4 e.kkrwlkneeiaaiLenfekheltl..elktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenpt 93
e ++rwl++ e+++iL+n+ k+ ++l ++ rp sgs++L++rk +ryfrkDG++w+kkkdgktvrE+he+LK g+v+vl+cyYah+e+n++
Pp3c8_2690V3.3.p 23 EaQTRWLRPVEVCEILKNYVKYGFKLnpVPPVRPCSGSMFLFDRKTLRYFRKDGHNWRKKKDGKTVREAHERLKTGSVDVLHCYYAHGEDNSS 115
4499*****************9998866899************************************************************** PP
CG-1 94 fqrrcywlLeeelekivlvhylev 117
fqrrcyw+L+ +le+ivlvhy+ev
Pp3c8_2690V3.3.p 116 FQRRCYWMLDPSLEHIVLVHYREV 139
**********************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 81.461 | 17 | 145 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 7.6E-72 | 20 | 140 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.7E-50 | 23 | 138 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 2.9E-6 | 537 | 625 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 3.9E-8 | 538 | 624 | IPR002909 | IPT domain |
| CDD | cd00102 | 0.0076 | 538 | 625 | No hit | No description |
| SuperFamily | SSF81296 | 3.62E-17 | 539 | 625 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 2.02E-17 | 700 | 834 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.9E-17 | 719 | 835 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 2.6E-6 | 721 | 801 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 3.49E-15 | 723 | 828 | No hit | No description |
| PROSITE profile | PS50297 | 17.767 | 740 | 834 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.0021 | 773 | 802 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 9.404 | 773 | 805 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 4300 | 812 | 843 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 130 | 897 | 919 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 6.888 | 898 | 926 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 32 | 943 | 965 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.035 | 944 | 973 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.657 | 963 | 992 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0044 | 985 | 1007 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.45 | 986 | 1010 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0013 | 988 | 1007 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 67 | 1060 | 1082 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.766 | 1062 | 1090 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1141 aa Download sequence Send to blast |
MAASRSRSGS QQPDIDIRQI ISEAQTRWLR PVEVCEILKN YVKYGFKLNP VPPVRPCSGS 60 MFLFDRKTLR YFRKDGHNWR KKKDGKTVRE AHERLKTGSV DVLHCYYAHG EDNSSFQRRC 120 YWMLDPSLEH IVLVHYREVT EGGRFTMSDS QHSAPAAHAA SPPEVSNPAT SPELDQDDAD 180 VIESEEVEEH SSLDPIVSPN APQDTLLDQL TPSARASADL NNSLMNGWIG GGSAASNMWM 240 PSSNRVGTSS LENLLNAPTG LTPRIGPFPQ QSSDKQQLGN NQFPDMFGTQ EPFSNGQRQE 300 SFTGAIDVAG WSRQMMDNNT RNNSTFGQSS DGPFAPHTAS GPPTYVKQES MGGATMWSDI 360 LEQCTTPSTH RNIMESLMNN NQLEMSDSPR TQDLGSSLVP PSKGILEALS PRGLGREPQT 420 PLEASLQAVT QEKALKSAGL YQRRLYGMRS ESQQLDLQFE RKSPMEKSPE SEPFPKLDSF 480 GRWMSHTLEA QNFPSPGAFQ SSWPGTPTMG FPGLPDDMYQ MQVDASLSTS VVADQRYSIT 540 DSSPEWAYAS EGAKVLVSGV FLGDYANASD IKWHCMFGEV EVPAEVINGV VLRCKAPPHA 600 PGRVSFYVTC GDRQAHSEIR EFEYRAGGRV VPDTKVERQA TEERLLKVRL AQLLLAESDE 660 DPGQLDDATD MKLQCLSPFS SSHSHSDWQE MENLANTSDH SQDINFHDRL LQMLLKSRMQ 720 KWLVCKIQEE GKGPSVLDAQ GQGVLHMAAA LGYDWAIAPM LAAGVPINFR DARGWTALHW 780 AASFAKEQVV IALLGHNADP GAVTDPTPSC PSGQTPADLA LMKGHVGIGG FLAESSLTRR 840 LNAMSLADQS PITVANAQFA GETAVAKLSR RDSIKHTLSS KSEDSVSIKE SLQAVRNAAR 900 AAALIQAAFR QHSFRKREEA EEEEASMKFD EYGMTESQMR GFVAARAAQR IQKAYRGHQE 960 KKQQLAASRI QQKFRSWKVR KDFVNLRQRV VRIQAHVRGN LVRRRFKKLL WSVGVLDKVI 1020 LRWRRKRSGL RGFKSGDLGI DTKEDDEEFL KEGRKLAEKA VEKAVTTVQT MVRSQPARDQ 1080 YRRLLEGTIK AEKLGQFQQA SPQEFHTSPD VEDYSYQELQ NQHMNQYLPH NGDQHMTFTD 1140 * |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Ppa.16890 | 0.0 | protonema | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
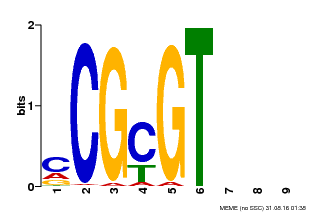
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024381994.1 | 0.0 | calmodulin-binding transcription activator 3-like | ||||
| Refseq | XP_024381996.1 | 0.0 | calmodulin-binding transcription activator 3-like | ||||
| Refseq | XP_024381997.1 | 0.0 | calmodulin-binding transcription activator 3-like | ||||
| TrEMBL | A0A2K1K5Y1 | 0.0 | A0A2K1K5Y1_PHYPA; Uncharacterized protein | ||||
| STRING | PP1S35_152V6.1 | 0.0 | (Physcomitrella patens) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 1e-122 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pp3c8_2690V3.3.p |



