 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pp3c2_23660V3.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Bryophytina; Bryopsida; Funariidae; Funariales; Funariaceae; Physcomitrella
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 940aa MW: 102762 Da PI: 7.1302 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 43.5 | 8.1e-14 | 247 | 295 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s+++GV ++grW A+I++ ++r++lg+f t+eeAa+a++ a+ k++g
Pp3c2_23660V3.1.p 247 SQFRGVV-PQSNGRWGAQIYEK-----HQRIWLGTFNTEEEAARAYDTAAIKFRG 295
6899996.6779*********3.....5************************998 PP
| |||||||
| 2 | B3 | 98.6 | 3.8e-31 | 412 | 512 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-S CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldg 86
f+k+ tpsdv+k++rlv+pk++ae++ +++ ++tl++ed sg+ W+++++y+++s++yvltkGW++Fvk+++L++gD+v F++
Pp3c2_23660V3.1.p 412 FDKAVTPSDVGKLNRLVIPKQHAERCfpldlsA-NSP-GQTLSFEDVSGKHWRFRYSYWNSSQSYVLTKGWSRFVKEKKLDAGDIVSFERG- 500
89***************************9853.433.459************************************************43. PP
SSEE..EEEEE-S CS
B3 87 rsefelvvkvfrk 99
++ el++ ++rk
Pp3c2_23660V3.1.p 501 -RNHELYIDFRRK 512
.667779999987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 1.18E-17 | 247 | 303 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 1.21E-25 | 247 | 305 | No hit | No description |
| SMART | SM00380 | 1.1E-30 | 248 | 309 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 20.521 | 248 | 303 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 2.5E-21 | 248 | 303 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 9.4E-5 | 249 | 260 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 7.7E-8 | 249 | 295 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 9.4E-5 | 269 | 285 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 7.6E-42 | 403 | 513 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.06E-30 | 408 | 512 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 6.77E-27 | 410 | 511 | No hit | No description |
| SMART | SM01019 | 1.4E-28 | 412 | 513 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 15.679 | 412 | 513 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 6.7E-28 | 412 | 512 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0048527 | Biological Process | lateral root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 940 aa Download sequence Send to blast |
MWSKPASRSS SEPHHGPDVT LRLNNFDTLV ASSSDSDVSS FKIVECPSQG REEASTSVVI 60 RETNPASPIP KFYTIEEKTA MLGVAEAAAA NTLQVHGIPR FQRHIDHDRS RLHDSSMLRT 120 SGKDSENCFR ADAEEHRAYV ASNLESFGER SHPLDLLGSR LEAEFSRGAN LKQWPLGSRN 180 VGQRCGSGEL ERTQSGPSAF LGSGSLDTEQ RGSLERTHSG PSFSGSGDSE YGDDRGREQS 240 NSKLPSSQFR GVVPQSNGRW GAQIYEKHQR IWLGTFNTEE EAARAYDTAA IKFRGRDAMT 300 NFRPVTDSEY ESEFLRSFSK EQIVEMLRRH TYDEELDQCK KVFNMDAAAN PVSARRRVLE 360 MCRSSNPGTR LSVGSFSLLQ NTFSADTSTT LAPPNLPRDE PRESSPTREH LFDKAVTPSD 420 VGKLNRLVIP KQHAERCFPL DLSANSPGQT LSFEDVSGKH WRFRYSYWNS SQSYVLTKGW 480 SRFVKEKKLD AGDIVSFERG RNHELYIDFR RKQITAGGTS TSDRSSFRSA WSGPCNNAYA 540 PSNGPVSPLN SRMWQPFSFS VPHVTVTSSG IQPSSMYGSI YNQMMQSARG TSAASGDLGS 600 VQDSAIPSMA GSCLNLDLQH SPLKCHAATE DFILHIDRPL RQSADMDGGS NIHDRSLVAS 660 SSSTSSQSEE APSTSSGTRL FGVDLERAAP LACKVSPLQS IPSFTSSALQ SKLSEGHESS 720 QGNLKTIHDG GNIPPSASLS ALTSGEFLSF RLGNAFKSLQ CKHPVEQSSP ENGGAPVRLS 780 LQASTSSSSG HRGSSPLKST AMTEGEDPKA RSSTESVASA LDSNSSRSSP MLLEKRKREN 840 SNRDYMQGCS DQADGAANVP LTLAHSEFRS SLEQARHAWH KHEDDASAEV QTSVKRHCSP 900 IQASLLRQGS GTTEQSDFLP KPDLVQEKQK PQYDSSNLR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5os9_A | 2e-51 | 408 | 512 | 4 | 113 | B3 domain-containing transcription factor NGA1 |
| 5os9_B | 2e-51 | 408 | 512 | 4 | 113 | B3 domain-containing transcription factor NGA1 |
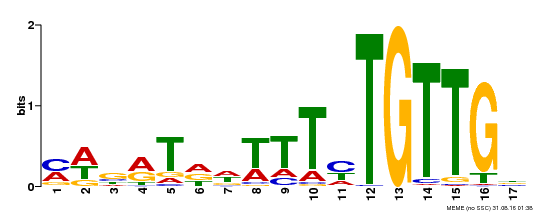
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00139 | DAP | Transfer from AT1G13260 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024369138.1 | 0.0 | uncharacterized protein LOC112279179 isoform X2 | ||||
| TrEMBL | A0A2K1L2S1 | 0.0 | A0A2K1L2S1_PHYPA; Uncharacterized protein | ||||
| STRING | PP1S135_69V6.1 | 0.0 | (Physcomitrella patens) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G13260.1 | 4e-90 | related to ABI3/VP1 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pp3c2_23660V3.1.p |



