 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Potri.006G020600.1 | ||||||||
| Common Name | POPTR_0006s02140g | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Populus
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 843aa MW: 97071 Da PI: 7.1243 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 89.8 | 3.6e-28 | 77 | 164 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHelap 91
+fY+eYAk++GF++++++s++sk+++e+++++f Cs++g + e+++ ++r++++++t+Cka+++vk++ dgkw ++++++eHnHel p
Potri.006G020600.1 77 SFYQEYAKSMGFTTSIKNSRRSKKSKEFIDAKFACSRYGVTPESDSG---NSRRSTVKKTDCKASMHVKRRADGKWIIHEFVKEHNHELLP 164
5****************************************999888...888999********************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 1.4E-25 | 77 | 164 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 1.0E-23 | 285 | 377 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.099 | 564 | 600 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 5.8E-4 | 574 | 598 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 2.5E-5 | 575 | 602 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 843 aa Download sequence Send to blast |
MMTERNHKMI DDMIDLQDNV PADDVVGGNI VGVVDVVHSR DVAVVDSPKR AVAMFEGDVN 60 YELCDGIEFG SHEEAYSFYQ EYAKSMGFTT SIKNSRRSKK SKEFIDAKFA CSRYGVTPES 120 DSGNSRRSTV KKTDCKASMH VKRRADGKWI IHEFVKEHNH ELLPALAYHF RIHRNVKLAE 180 KNNIDILHAV SERTRKMYVE MSRQSGGYQN FGLVKSEMNM QFEKGQHLAL DEGDAQVVLE 240 YFKRVKKENA NFFYAIDLNE EQRLRNLFWV DAKSRADYIS FNDAVCFETF YVKYHEKLPF 300 APFVGVNHHC QPILLGCAFI ADESRSTFVW LMKTWLRAMG GQAPKVIVTD VDKTLKVAIE 360 EVFPNTRHCF SLWHILERLP ETLSHVIKRH ENFLPKFNKC IFKSWTDDRF DMRWWKMVTR 420 FELQDDEWIQ SLYEDRKKWV PTYMGDTFLA GTSATQRSES MSAFFDKYIH RKITMKEFMK 480 QYGTILQNRY EDESVADFDT SHKQPALKSP SPWEKQMSMV YTHAIFKKFQ VEVLGVVGCH 540 PKKESEDGTL VTFRVQDCEK DEHFLVTWNQ TNSEVCCFCH SFEYKGFLCR HALIVLQICG 600 LSNIPPHYIL KRWTKDAKSR QPMAVGTERA QTRVQRYNDL CKLAIEMSEE GSLSEESYNI 660 VLHTLVEALK NCVNVNNCNN SVAESSTYTL THREAEEENQ GSLVTKSSKK KNPVRKRKVQ 720 SDPDVMLVEA PDSLQQMENL SSEGINLGGY YGTQQNVQGL VQLNLMEPPH DGYYVNQQSM 780 QGLGQLNSIA PSHDGFFGTQ QSLHGLGQYD FRPPTGFSYS MQLQDDTHLR SSHMHGSASR 840 HA* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
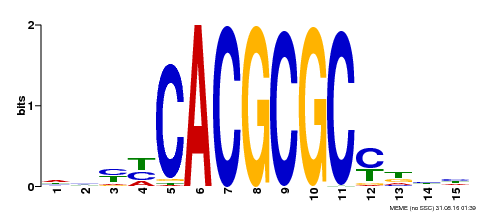
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Potri.006G020600.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024459852.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 isoform X1 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A2K1ZVH1 | 0.0 | A0A2K1ZVH1_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0006s02140.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF40 | 30 | 530 |
| Representative plant | OGRP13457 | 3 | 4 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Potri.006G020600.1 |
| Entrez Gene | 7495635 |



