 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Phvul.008G270400.2 | ||||||||
| Common Name | PHAVU_008G270400g | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Phaseolus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 320aa MW: 37064.5 Da PI: 8.355 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 20.5 | 1.2e-06 | 63 | 85 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C+dCg +F+ + +L +H+++H
Phvul.008G270400.2 63 HTCEDCGITFKKHAYLLQHMQSH 85
79*******************99 PP
| |||||||
| 2 | zf-C2H2 | 19 | 3.9e-06 | 91 | 115 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C dC s++rk++L+rH+ +H
Phvul.008G270400.2 91 FVCAldDCQASYRRKDHLTRHLLLH 115
89********************998 PP
| |||||||
| 3 | zf-C2H2 | 15.1 | 6.6e-05 | 120 | 145 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
+kCp +C++ Fs sn+krH H
Phvul.008G270400.2 120 FKCPvvNCNTEFSLLSNMKRHVEEiH 145
89*******************98777 PP
| |||||||
| 4 | zf-C2H2 | 20 | 1.8e-06 | 160 | 184 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++Cp Cgk+F+ s+Lk+H +H
Phvul.008G270400.2 160 HVCPeiGCGKVFRFASQLKKHEDSH 184
79*******************9888 PP
| |||||||
| 5 | zf-C2H2 | 16.4 | 2.6e-05 | 251 | 276 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
++C+ C+ +Fs+ksnL +H ++ H
Phvul.008G270400.2 251 FQCEfkGCSCTFSNKSNLDKHKKVvH 276
89********************9988 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 28 | 19 | 41 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 8.725 | 19 | 46 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 7.8E-8 | 61 | 85 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.007 | 63 | 85 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.297 | 63 | 90 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 65 | 85 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.21E-10 | 71 | 124 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.0E-11 | 86 | 117 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.049 | 91 | 115 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.097 | 91 | 120 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 93 | 115 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.2E-5 | 118 | 146 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.2E-5 | 118 | 146 | No hit | No description |
| SMART | SM00355 | 0.017 | 120 | 145 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.949 | 120 | 150 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 122 | 145 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.6E-7 | 156 | 184 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 8.96E-5 | 157 | 186 | No hit | No description |
| SMART | SM00355 | 0.0051 | 160 | 184 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.408 | 160 | 189 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 162 | 184 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 2.6 | 192 | 217 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 194 | 217 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 35 | 220 | 241 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 7.08E-9 | 233 | 286 | No hit | No description |
| PROSITE profile | PS50157 | 13.547 | 251 | 281 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0029 | 251 | 276 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.8E-6 | 251 | 275 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 253 | 276 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.4E-5 | 276 | 305 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 12 | 282 | 308 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 284 | 308 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 320 aa Download sequence Send to blast |
MVEAVEIEER PKFRDIRRYF CQYCGICRSK KTLITSHINS HHKEEDEKAR KERDPEEESS 60 KSHTCEDCGI TFKKHAYLLQ HMQSHSLERP FVCALDDCQA SYRRKDHLTR HLLLHEGKTF 120 KCPVVNCNTE FSLLSNMKRH VEEIHDDSSP AVTDKSHKQH VCPEIGCGKV FRFASQLKKH 180 EDSHVKLESV DVVCLEPGCM MHFTNVQCLK AHVNSSHQYM TCDICGSKQL KKNIKRHLRS 240 HEPDSSSLET FQCEFKGCSC TFSNKSNLDK HKKVVHLKEK PFLCGFPDCG MRFAYKHVRD 300 KHEKTVKHVF TLVSSTFLL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1tf6_A | 8e-15 | 67 | 212 | 19 | 155 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| 1tf6_D | 8e-15 | 67 | 212 | 19 | 155 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
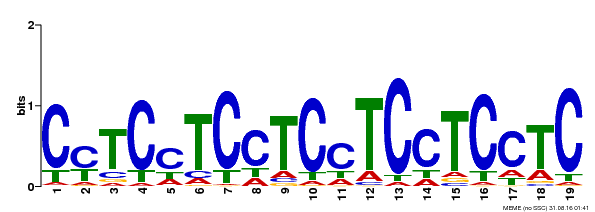
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Phvul.008G270400.2 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015036 | 1e-111 | AP015036.1 Vigna angularis var. angularis DNA, chromosome 3, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007142320.1 | 0.0 | hypothetical protein PHAVU_008G270400g | ||||
| Swissprot | Q84MZ4 | 1e-121 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | V7B924 | 0.0 | V7B924_PHAVU; Uncharacterized protein | ||||
| STRING | XP_007142319.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-119 | transcription factor IIIA | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Phvul.008G270400.2 |
| Entrez Gene | 18624175 |



