 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Peinf101Scf02920g02003.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Petunioideae; Petunia; Petunia integrifolia
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 468aa MW: 51070.8 Da PI: 5.0597 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 52.7 | 1e-16 | 55 | 99 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT+eE++++v+a k++G W+ I +++g ++t+ q++s+ qk+
Peinf101Scf02920g02003.1 55 REKWTEEEHQRFVEALKLYGRS-WRQIEEHVG-TKTAIQIRSHAQKF 99
789*****************55.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 4.71E-16 | 49 | 103 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 22.393 | 50 | 104 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.4E-16 | 53 | 102 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 3.8E-13 | 54 | 102 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.2E-9 | 54 | 95 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 9.7E-14 | 55 | 98 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.49E-10 | 57 | 100 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 468 aa Download sequence Send to blast |
MVAEEMNQIE GARQGASSMI SSKLETVAAQ GKALNCLANE NMLKVRKPYT ITKQREKWTE 60 EEHQRFVEAL KLYGRSWRQI EEHVGTKTAI QIRSHAQKFF AKVARDSGID GDESLNAIDI 120 PPPRPKKKPL HPYPRKMVDS PVADKAVSGQ PESSPSPNAS GRDGRSPDSV LSAIGSDVFE 180 YAVVEQQPSR FSPASCTTDA QTANIISAEN DESSMTSNSS TVDKIHVASK PVAPNTSLIT 240 NYGMECDLFP RGGSCSGESQ AVEAPLASIK LFGQTVLVPD ATKLTLGAPN SCNSLPSIAT 300 EKEINNENLF HGFQGNQANS QFIVAMVPGN IIPPDCWPSQ YRLENNLEGV APFPDTVSWW 360 SWYQDLLYRY ISSCSQTAME TAAPCQASKD EEPQREGSST GSSVGSASEV DDGNRSSETA 420 ESKCTTKSDK WNNIKGFVPY KRCLAERDEN SSGVVLEERE SQRVRVCS |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00146 | DAP | Transfer from AT1G18330 | Download |
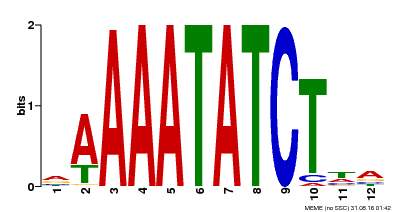
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019224849.1 | 0.0 | PREDICTED: protein REVEILLE 1-like isoform X1 | ||||
| TrEMBL | A0A314KU97 | 0.0 | A0A314KU97_NICAT; Protein reveille 7 | ||||
| STRING | XP_009768903.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA7447 | 21 | 27 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G18330.1 | 3e-41 | MYB_related family protein | ||||



