 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Peinf101Scf00889g04016.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Petunioideae; Petunia; Petunia integrifolia
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 357aa MW: 38881.5 Da PI: 8.8357 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 129.2 | 1.5e-40 | 50 | 126 | 2 | 78 |
-SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 2 CqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
C v+gC+adls+++eyhrrhkvCevhsk+++v + g+eqrfCqqCsrfh+l efD++krsCr+rL++hn+rrrk+q+
Peinf101Scf00889g04016.1 50 CLVDGCNADLSQCREYHRRHKVCEVHSKTAKVSIGGREQRFCQQCSRFHSLVEFDDGKRSCRKRLDGHNRRRRKPQP 126
**************************************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.4E-32 | 45 | 111 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.744 | 47 | 124 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 4.45E-38 | 48 | 129 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 4.5E-30 | 50 | 123 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 357 aa Download sequence Send to blast |
IVLDFKEYKL KPLLNLAIGR FLLYENCVME SSGSSKRAKA PGSLAQVAHC LVDGCNADLS 60 QCREYHRRHK VCEVHSKTAK VSIGGREQRF CQQCSRFHSL VEFDDGKRSC RKRLDGHNRR 120 RRKPQPDPMA KNSALLFSGQ QGTKLLSFGS QQIFPSAVVS SAWAGVVKTD SEMVLYNNQS 180 HINCIDSQNS FPDSSAHSSY KGGSQFQFIQ GSDHSSLPES SICQPLHDPP NSAAGIPSSG 240 RKIFSSGLNE IVDSDRALSL LSSAPAVTRE IGFSHMVQQP AASITRPQSH AAVHGLHYGG 300 LSQYPYAQDF NSKPQNSAAN SHVSNSSPLH FPGMLQNAPD GSSTSGGSQQ TLAFMWD |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 4e-28 | 50 | 123 | 11 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 106 | 123 | KRSCRKRLDGHNRRRRKP |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00555 | DAP | Transfer from AT5G50570 | Download |
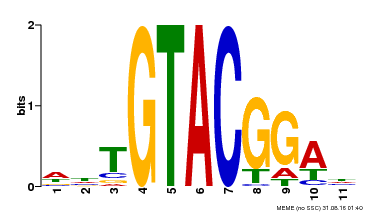
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LC073300 | 0.0 | LC073300.1 Nicotiana tabacum NtabSPL13 mRNA for squamosa promoter binding protein NtabSPL13, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001311938.1 | 0.0 | teosinte glume architecture 1-like | ||||
| Refseq | XP_009621035.1 | 0.0 | PREDICTED: teosinte glume architecture 1-like | ||||
| Refseq | XP_009621036.1 | 0.0 | PREDICTED: teosinte glume architecture 1-like | ||||
| Refseq | XP_009621037.1 | 0.0 | PREDICTED: teosinte glume architecture 1-like | ||||
| Refseq | XP_009621038.1 | 0.0 | PREDICTED: teosinte glume architecture 1-like | ||||
| Refseq | XP_016443564.1 | 0.0 | PREDICTED: teosinte glume architecture 1-like isoform X1 | ||||
| Refseq | XP_016443565.1 | 0.0 | PREDICTED: teosinte glume architecture 1-like isoform X1 | ||||
| Refseq | XP_016443566.1 | 0.0 | PREDICTED: teosinte glume architecture 1-like isoform X1 | ||||
| Refseq | XP_016443567.1 | 0.0 | PREDICTED: teosinte glume architecture 1-like isoform X1 | ||||
| TrEMBL | A0A2K9ZXK2 | 0.0 | A0A2K9ZXK2_PETHY; Squamosa promoter binding-like protein 13 | ||||
| STRING | XP_009621034.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4920 | 20 | 37 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G50670.1 | 3e-48 | SBP family protein | ||||



