 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Peinf101Scf00331g03006.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Petunioideae; Petunia; Petunia integrifolia
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 527aa MW: 58115.9 Da PI: 6.9557 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 42.5 | 1.2e-13 | 353 | 393 | 10 | 55 |
HHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 10 rrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+rRRdriN+++ L+ellP++ +K +Ka++L +A+eY+ksLq
Peinf101Scf00331g03006.1 353 QRRRDRINEKMKALQELLPHS-----TKTDKASMLDEAIEYLKSLQ 393
79******************9.....9******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00083 | 3.31E-14 | 338 | 397 | No hit | No description |
| PROSITE profile | PS50888 | 14.5 | 343 | 392 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.9E-12 | 349 | 398 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 6.3E-11 | 353 | 393 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 1.7E-15 | 353 | 417 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 8.8E-16 | 353 | 401 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009704 | Biological Process | de-etiolation | ||||
| GO:0010161 | Biological Process | red light signaling pathway | ||||
| GO:0010244 | Biological Process | response to low fluence blue light stimulus by blue low-fluence system | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0010928 | Biological Process | regulation of auxin mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 527 aa Download sequence Send to blast |
MDGLFYSFDH ELVELLWRNG EVVLHSQTHK KPGYDPNDCR LVNKHDQSTI RTAVSCGNQT 60 NLIQDDETIS WLNCPIDDPF DREFCSPFLS EISTNPIEAD KITRQSEDNK VFKFDPFEVN 120 HVLPHSHHSG FDPNPMPPPR YHNSGSAQQQ HVGGLQKGVN FPTPVRSSNV QLGGKEVSNM 180 MLHDVKEGSV MTVGSSHCGS NQVANDIDTS RFSSSTNRGL STAMITDYTG KVSPQSDTMD 240 RDTFEPANTS SSGGSGSSYA RTCNQSTATN SHSHKRKSRE GEEPECQSKS GDLDSAGGNK 300 SAQKSGTARR SRAAEVHNLS ERVEFVGLTV IYIHVMASCK EKHLNHQMAL SLQRRRDRIN 360 EKMKALQELL PHSTKTDKAS MLDEAIEYLK SLQMQLQMMW MGGGMASMMF PGVQHYMSRM 420 GIGMGPPTVS SIHNAMNLAR LPLVDPPIPL PQAAPNQAAM CQTSMLNAVN FQRHLQNPNF 480 SDQYASYMGF HPLQGASQPI NIFSLGSHSA QQSQQIPPPT NSTAPTT |
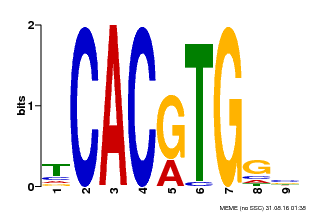
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00606 | ChIP-seq | Transfer from AT2G43010 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT013582 | 0.0 | BT013582.1 Lycopersicon esculentum clone 132330F, mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016580439.1 | 0.0 | PREDICTED: transcription factor PIF4 | ||||
| TrEMBL | A0A1U8HIU3 | 0.0 | A0A1U8HIU3_CAPAN; Uncharacterized protein | ||||
| TrEMBL | A0A2G2Z1M5 | 0.0 | A0A2G2Z1M5_CAPAN; transcription factor PIF4 | ||||
| STRING | Solyc07g043580.2.1 | 0.0 | (Solanum lycopersicum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA10465 | 22 | 26 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43010.2 | 2e-38 | phytochrome interacting factor 4 | ||||



