 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Peinf101Scf00181g08015.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Petunioideae; Petunia; Petunia integrifolia
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 196aa MW: 21968 Da PI: 10.3757 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.5 | 2.6e-32 | 128 | 181 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
pr+rWt++LH++Fv+av+ LGG+e+AtPk++lelm+vk+Ltl+hvkSHLQ+YR+
Peinf101Scf00181g08015.1 128 PRMRWTSTLHAHFVHAVQLLGGHERATPKSVLELMNVKDLTLAHVKSHLQMYRT 181
9****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 9.85E-16 | 126 | 182 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 9.9E-29 | 126 | 182 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.2E-24 | 128 | 181 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.3E-7 | 129 | 180 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005618 | Cellular Component | cell wall | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 196 aa Download sequence Send to blast |
MTEASSTMPD LSLQISLPSN IPTCDKFKQV LGFNGFTKSS SIVNNHSSDS GSSGSDLSQE 60 NGLFQPEKSF NVSLIPSIEP TLSLGLAQLP SRNLHHQQYQ HFQPQIYGRD FKRNSRMISG 120 VKRSCRAPRM RWTSTLHAHF VHAVQLLGGH ERATPKSVLE LMNVKDLTLA HVKSHLQMYR 180 TVKSTDKSTG IWNFLF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 5e-18 | 129 | 183 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 5e-18 | 129 | 183 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 5e-18 | 129 | 183 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 5e-18 | 129 | 183 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 5e-18 | 129 | 183 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 5e-18 | 129 | 183 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 5e-18 | 129 | 183 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 5e-18 | 129 | 183 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that regulates carpel integuments formation. Required for the specification of polarity in the ovule inner integument. Modulates the content of flavonols and proanthocyanidin in seeds. {ECO:0000269|PubMed:16623911, ECO:0000269|PubMed:19054366, ECO:0000269|PubMed:20444210}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00107 | PBM | Transfer from AT5G42630 | Download |
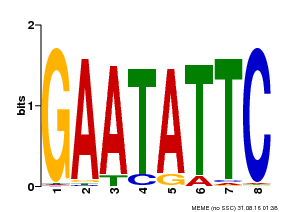
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KP271989 | 4e-72 | KP271989.1 UNVERIFIED: Populus trichocarpa isolate Potri.014G037200_BESC-125 mRNA sequence. | |||
| GenBank | KP271990 | 4e-72 | KP271990.1 UNVERIFIED: Populus trichocarpa isolate Potri.014G037200_BESC-293 mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006354964.2 | 4e-80 | PREDICTED: probable transcription factor KAN4 | ||||
| Swissprot | Q9FJV5 | 1e-53 | KAN4_ARATH; Probable transcription factor KAN4 | ||||
| TrEMBL | A0A2G3CQ52 | 4e-80 | A0A2G3CQ52_CAPCH; Putative transcription factor KAN4 | ||||
| STRING | PGSC0003DMT400030518 | 3e-78 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA12262 | 18 | 22 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G42630.2 | 7e-56 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



