 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Peaxi162Scf00503g00427.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Petunioideae; Petunia
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 334aa MW: 38125.8 Da PI: 6.6754 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 55.7 | 8.2e-18 | 122 | 175 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe +++ e++ +LA+ lgL+ rq+ +WFqNrRa++k
Peaxi162Scf00503g00427.1 122 KKRRLNMEQVKTLEKNFELGNKLEPERKMQLARALGLQPRQIAIWFQNRRARWK 175
456899***********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 131.7 | 2.8e-42 | 121 | 213 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeve 85
ekkrrl+ eqvk+LE++Fe +kLeperK++lar+Lglqprq+a+WFqnrRAR+ktkqlEkdy++Lkr++da+k+en++L+++++
Peaxi162Scf00503g00427.1 121 EKKRRLNMEQVKTLEKNFELGNKLEPERKMQLARALGLQPRQIAIWFQNRRARWKTKQLEKDYDVLKRQFDAIKAENDALQAQNQ 205
69*********************************************************************************** PP
HD-ZIP_I/II 86 eLreelke 93
+L++e+++
Peaxi162Scf00503g00427.1 206 KLHAEIMA 213
***99975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.75E-19 | 108 | 179 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.329 | 117 | 177 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.8E-17 | 120 | 181 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 3.6E-15 | 122 | 175 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.95E-15 | 122 | 178 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 4.9E-19 | 124 | 184 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 1.4E-5 | 148 | 157 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 152 | 175 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.4E-5 | 157 | 173 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 5.4E-17 | 177 | 219 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009744 | Biological Process | response to sucrose | ||||
| GO:0048826 | Biological Process | cotyledon morphogenesis | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 334 aa Download sequence Send to blast |
MHTYNKPKLH KLGKASLEAD KHESQSYIFG PVAKAFKVMT CTNEMAFFPT NFMLQTTHED 60 DQHQPSTSLT PILPSCSPQD FHGIASFLGK RSMSFSGMEA NNCEENHGED DLSDDGSQAG 120 EKKRRLNMEQ VKTLEKNFEL GNKLEPERKM QLARALGLQP RQIAIWFQNR RARWKTKQLE 180 KDYDVLKRQF DAIKAENDAL QAQNQKLHAE IMALKNMEQP TESINLNKET EGSCSNRSEN 240 SSEIKLDISR TPAIDSPLSN HPITSRPFFP PSMRPITNGV AQLFHNSNTS SRPDLQQCLK 300 NMDQSVKEES LSNMFCGIDD QTAFWPWLEQ QQFN |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 169 | 177 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may act in the sucrose-signaling pathway. {ECO:0000269|PubMed:11292072}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
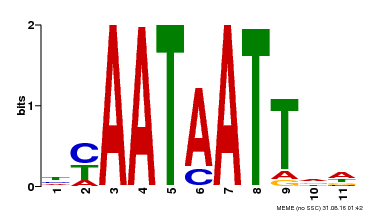
| Motif ID | Method | Source | Motif file |
| MP00225 | DAP | Transfer from AT1G69780 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC142505 | 1e-151 | AC142505.1 Solanum demissum chromosome 5 clone PGEC287I17, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016548012.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-13 | ||||
| Swissprot | Q8LC03 | 1e-125 | ATB13_ARATH; Homeobox-leucine zipper protein ATHB-13 | ||||
| TrEMBL | A0A2G2VLY1 | 0.0 | A0A2G2VLY1_CAPBA; Homeobox-leucine zipper protein HOX21 | ||||
| STRING | XP_009593379.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA848 | 24 | 98 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69780.1 | 1e-128 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



