 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Peaxi162Scf00136g00115.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Petunioideae; Petunia
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 303aa MW: 33620.1 Da PI: 9.4803 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 44 | 5.2e-14 | 118 | 162 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ +++ + ++lG+g+W+ I+r + ++Rt+ q+ s+ qky
Peaxi162Scf00136g00115.1 118 PWTEEEHRIFLLGLEKLGKGDWRGISRNFVTTRTPTQVASHAQKY 162
8*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.60.10 | 8.6E-4 | 3 | 20 | IPR001878 | Zinc finger, CCHC-type |
| PROSITE profile | PS51294 | 17.838 | 111 | 167 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 9.72E-18 | 113 | 168 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.1E-18 | 114 | 166 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 2.6E-11 | 115 | 165 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-11 | 115 | 161 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 2.0E-11 | 118 | 162 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 8.50E-10 | 118 | 163 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 303 aa Download sequence Send to blast |
MGRKCSHCGI TGHNSRTCKT QNLTKVDEET EDATTYNIGL KLFGVKLEIP KSSSSCVSMK 60 KSFSVGCLSK ISSSTTGSFV SNINCQMFMD HENMDKVSNG YLSDVLVGRT SDKKKGVPWT 120 EEEHRIFLLG LEKLGKGDWR GISRNFVTTR TPTQVASHAQ KYFLRQNSLN RKKRRRSLFD 180 EVGMDKSTIL LFDKSVTTKT NGTCITNISY EKSPDDETMI SMINFSTSLS KDSKSSTSQH 240 SMPATTLVNL PFINSSNNFT CHNISVQDSA KPDLELTLAV SKPLNNQRRQ PGCLNAENSI 300 AVI |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription repressor that binds to 5'-TATCCA-3' elements in gene promoters. Contributes to the sugar-repressed transcription of promoters containing SRS or 5'-TATCCA-3' elements. Transcription repressor involved in a cold stress response pathway that confers cold tolerance. Suppresses the DREB1-dependent signaling pathway under prolonged cold stress. DREB1 responds quickly and transiently while MYBS3 responds slowly to cold stress. They may act sequentially and complementarily for adaptation to short- and long-term cold stress (PubMed:20130099). {ECO:0000269|PubMed:12172034, ECO:0000269|PubMed:20130099}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00565 | DAP | Transfer from AT5G56840 | Download |
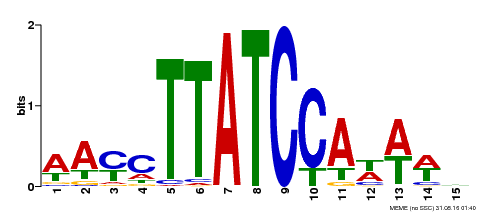
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Repressed by sucrose and gibberellic acid (GA) (PubMed:12172034). Induced by cold stress in roots and shoots. Induced by salt stress in shoots. Down-regulated by abscisic aci (ABA) in shoots (PubMed:20130099). {ECO:0000269|PubMed:12172034, ECO:0000269|PubMed:20130099}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP010894 | 2e-62 | AP010894.1 Solanum lycopersicum DNA, chromosome 8, clone: C08HBa0017M21, complete sequence. | |||
| GenBank | HG975520 | 2e-62 | HG975520.1 Solanum lycopersicum chromosome ch08, complete genome. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009631208.1 | 1e-138 | PREDICTED: myb-like protein J isoform X1 | ||||
| Refseq | XP_009780709.1 | 1e-138 | PREDICTED: myb-like protein H isoform X1 | ||||
| Refseq | XP_016450793.1 | 1e-138 | PREDICTED: myb-like protein H isoform X1 | ||||
| Refseq | XP_016498568.1 | 1e-138 | PREDICTED: myb-like protein J | ||||
| Swissprot | Q7XC57 | 1e-50 | MYBS3_ORYSJ; Transcription factor MYBS3 | ||||
| TrEMBL | A0A1S3YFJ3 | 1e-137 | A0A1S3YFJ3_TOBAC; myb-like protein H isoform X1 | ||||
| TrEMBL | A0A1S4CBQ0 | 1e-137 | A0A1S4CBQ0_TOBAC; myb-like protein J | ||||
| TrEMBL | A0A1U7X1F4 | 1e-137 | A0A1U7X1F4_NICSY; myb-like protein H isoform X1 | ||||
| STRING | XP_009780709.1 | 1e-138 | (Nicotiana sylvestris) | ||||
| STRING | XP_009631208.1 | 1e-138 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3385 | 21 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G47390.1 | 2e-47 | MYB_related family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



