 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Peaxi162Scf00089g00844.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Petunioideae; Petunia
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1227aa MW: 137857 Da PI: 8.9816 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.8 | 0.00037 | 1112 | 1137 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
y+C C++sFs+k +L H ++ +
Peaxi162Scf00089g00844.1 1112 YQCDleGCSMSFSSKQELLLHKKNvC 1137
99*******************99866 PP
| |||||||
| 2 | zf-C2H2 | 11 | 0.0014 | 1136 | 1159 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirtH 23
+Cp C+k F ++ +L++H r+H
Peaxi162Scf00089g00844.1 1136 VCPveGCKKKFFSHKYLVQHRRVH 1159
69999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 12.2 | 0.00054 | 1195 | 1221 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C+ Cg++F+ s++ rH r+ H
Peaxi162Scf00089g00844.1 1195 YVCTetGCGQTFRFVSDFSRHKRKtgH 1221
99********************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 1.1E-14 | 17 | 58 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 13.384 | 18 | 59 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 8.7E-15 | 19 | 52 | IPR003349 | JmjN domain |
| PROSITE profile | PS51184 | 21.97 | 186 | 322 | IPR003347 | JmjC domain |
| SMART | SM00558 | 1.1E-32 | 186 | 322 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 1.76E-18 | 200 | 338 | No hit | No description |
| Pfam | PF02373 | 2.7E-5 | 219 | 247 | IPR003347 | JmjC domain |
| Pfam | PF02373 | 1.1E-16 | 250 | 305 | IPR003347 | JmjC domain |
| SMART | SM00355 | 7.4 | 1112 | 1134 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 5.0E-5 | 1133 | 1163 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.012 | 1135 | 1159 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.572 | 1135 | 1164 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1137 | 1159 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 3.02E-9 | 1151 | 1191 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.8E-8 | 1164 | 1189 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0014 | 1165 | 1189 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.741 | 1165 | 1194 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1167 | 1189 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 5.55E-8 | 1183 | 1217 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.5E-9 | 1190 | 1218 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.19 | 1195 | 1221 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.053 | 1195 | 1226 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1197 | 1221 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1227 aa Download sequence Send to blast |
MAEVNIEVFP WLKTLPVAPE YHPTLEEFQD PIAYIFKIEK HASQFGICKI VPPVPPLAKK 60 TILSNLNRSL SARPGSTGPT FTTRQQQIGF CPRKHRPVKK PVWQSGETYT VAEFQGKAKN 120 FEKNYLKKNC FFKKNAALSP LEIETLYWKA TVDKPFSVEY ANDMPGSAFP PKKIGGNGVG 180 GGEGVSLADT EWNMRGVSRA KGSLLKFMKE EIPGVTSPMV YLAMMFSWFA WHVEDHDLHS 240 LNYMHMVTFA TLGEKTTVMS PEVLLSAGIP CCRLVQNAGE FVVTFPRAYH SGFSHGFNCG 300 EAANIATPEW LRFAKEAAIR RASTNCPPMV SHFQLLYDLA LSLCSRVPKN IKIEPRSSRL 360 KDKKRSEGDM LVKELFVEDL NCNNYLLHIL GEGAPVVLLP QDYTGTSNSS YLVAGSQLKV 420 NSRSFPSLSS RDHEVKSLKD SASDDLMLGR KRGMKQLPRS SLEKGKYSSW HAGNRMPDSG 480 RNDDAQSSPE TKRGNLDTAR EMAYKCDTLS EHGLFSCVTC GILCYTCVAI VQPTEEAARH 540 LISSDFRNFN DWTGDVGGVT AIGRDLNAAD SDSSSGWLVK RSPGGLIDVP IESSDRIRKL 600 NNESVGVLSS TKARKETSSL GLLALNYANS SDSDEDEVEA DIPIEACESI HTDSEDEVSL 660 RVIDPYANHR QKRAVSEGRI CQRFDNSIHL ESEHSPSGES NTVSDRLRHH PRSHQVAANC 720 TPFSHKEEMN NNNDVAPFDN MPMQFTSTSD EDSFRMHVFC LQHAVQVEEQ LRQVGGVRIS 780 LLCHPDYPKL EAQAKKVAEE LGSDHFWREI SFREATKEDE EMIQSALEIE EAIHGNGDWT 840 VKLDMNLFYS ANLSRSPLYS KQMPYNFIIY SAFGRSSPDN TPEKSEYTGR GSGKQKRGVV 900 AGKWCGKVWM SSQVHPLLVE RDTDEEQEQN KSIPVQIKPD VKSERPSERA GTRTVATTCK 960 TGRKRKSAAE DRNTSNRKLL IADDLDDSLL SYIPQQHRKT NLRSKRIKYE TPEPQQDVDK 1020 KKRFSTPSDD ELDGGPSTRL RKRIPKPSKE SAAKLVKAKS AIKQHEGTKA EKGSKVKIPS 1080 TNRNTKKDPV MKAPRNNTGN TIKKMKDKEG EYQCDLEGCS MSFSSKQELL LHKKNVCPVE 1140 GCKKKFFSHK YLVQHRRVHM DDRPLKCPWK GCKMTFKWAW ARTEHIRVHT GARPYVCTET 1200 GCGQTFRFVS DFSRHKRKTG HVSKKGR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6a57_A | 2e-64 | 1112 | 1225 | 23 | 136 | Lysine-specific demethylase REF6 |
| 6a58_A | 2e-64 | 1112 | 1225 | 23 | 136 | Lysine-specific demethylase REF6 |
| 6a59_A | 2e-64 | 1112 | 1225 | 23 | 136 | Lysine-specific demethylase REF6 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 448 | 455 | GRKRGMKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
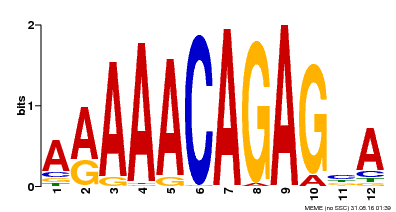
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006351452.1 | 0.0 | PREDICTED: lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | M1B7Y2 | 0.0 | M1B7Y2_SOLTU; Uncharacterized protein | ||||
| STRING | PGSC0003DMT400039224 | 0.0 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4988 | 23 | 31 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



