 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pbr030767.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Pyrus
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 373aa MW: 43092 Da PI: 6.8843 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 176.3 | 8.6e-55 | 8 | 138 | 1 | 129 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpk..kvka.eekewyfFskrdkkyatgkrknratksgyWkatgkdkevlsk 95
+ppGfrF+Ptdeelv++yL+kk++ k++++ +vi++vd+yk+ePwdL+ k+ + +++ewyfFs++dkky+tg+r+nratk+g+Wkatg+dk++++
Pbr030767.1 8 VPPGFRFRPTDEELVDYYLRKKIALKPIDF-DVIRDVDLYKIEPWDLEDlcKIGSgDQNEWYFFSHKDKKYPTGSRTNRATKAGFWKATGRDKAIYRG 104
69****************************.9***************943466553556**************************************9 PP
NAM 96 kgelvglkktLvfykgrapkgektdWvmheyrle 129
++ lvg++ktLvfykgrap+g+k+dW+mheyrle
Pbr030767.1 105 EKFLVGMRKTLVFYKGRAPNGQKSDWIMHEYRLE 138
9999****************************85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 5.36E-60 | 5 | 158 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 59.243 | 8 | 158 | IPR003441 | NAC domain |
| Pfam | PF02365 | 4.0E-28 | 9 | 137 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0048759 | Biological Process | xylem vessel member cell differentiation | ||||
| GO:1901348 | Biological Process | positive regulation of secondary cell wall biogenesis | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 373 aa Download sequence Send to blast |
MNTLSSHVPP GFRFRPTDEE LVDYYLRKKI ALKPIDFDVI RDVDLYKIEP WDLEDLCKIG 60 SGDQNEWYFF SHKDKKYPTG SRTNRATKAG FWKATGRDKA IYRGEKFLVG MRKTLVFYKG 120 RAPNGQKSDW IMHEYRLETN ENGTPQAKGW VVCRVFKKKL AAVSRMGDYE SPDCYDQVSF 180 LPQLDNPPRT SMSRTYASQN EQQLHYSQCK QEFDLLYNMP HHHHDSFLQL PQLESPKVPH 240 QSATPYGNCV MQSSTLTQEE QFAQYISQQN LFKSSSSPSV HALDNNNDYL QAVDQVKDWR 300 ALDKYVASQL SHDQQDASKE VTNYSHAEEI FHVAEHINML ANDASRRPDN NNNIGQDYAS 360 TSTSTFQIDL WK* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 7e-47 | 8 | 162 | 17 | 169 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 7e-47 | 8 | 162 | 17 | 169 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 7e-47 | 8 | 162 | 17 | 169 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 7e-47 | 8 | 162 | 17 | 169 | NO APICAL MERISTEM PROTEIN |
| 4dul_A | 7e-47 | 8 | 162 | 17 | 169 | NAC domain-containing protein 19 |
| 4dul_B | 7e-47 | 8 | 162 | 17 | 169 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the secondary wall NAC binding element (SNBE), 5'-(T/A)NN(C/T)(T/C/G)TNNNNNNNA(A/C)GN(A/C/T)(A/T)-3', in the promoter of target genes (By similarity). Involved in xylem formation by promoting the expression of secondary wall-associated transcription factors and of genes involved in secondary wall biosynthesis and programmed cell death, genes driven by the secondary wall NAC binding element (SNBE). Triggers thickening of secondary walls (PubMed:16103214, PubMed:25148240). {ECO:0000250|UniProtKB:Q9LVA1, ECO:0000269|PubMed:16103214, ECO:0000269|PubMed:25148240}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00136 | DAP | Transfer from AT1G12260 | Download |
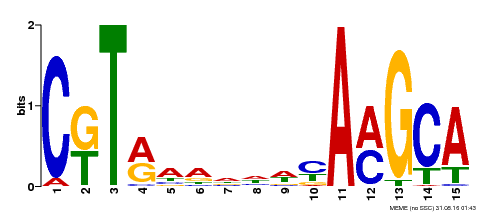
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pbr030767.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009339709.1 | 0.0 | PREDICTED: NAC domain-containing protein 7-like isoform X2 | ||||
| Refseq | XP_009339710.1 | 0.0 | PREDICTED: NAC domain-containing protein 7-like isoform X2 | ||||
| Refseq | XP_018499169.1 | 0.0 | PREDICTED: NAC domain-containing protein 7-like isoform X1 | ||||
| Swissprot | Q9FWX2 | 1e-128 | NAC7_ARATH; NAC domain-containing protein 7 | ||||
| TrEMBL | A0A498JZ27 | 0.0 | A0A498JZ27_MALDO; Uncharacterized protein | ||||
| STRING | XP_009339709.1 | 0.0 | (Pyrus x bretschneideri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1269 | 33 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G12260.1 | 1e-130 | NAC 007 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



