 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pbr028458.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Pyrus
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1078aa MW: 120187 Da PI: 5.3805 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 183.3 | 2.8e-57 | 20 | 136 | 2 | 118 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcy 99
++ ++rwl++ ei++iL nf+k++++ e++++p+sgsl L++rk++ryfrkDG++w+kk+dgktv+E+hekLKvg+v+vl+cyYah+e++++fqrrcy
Pbr028458.1 20 VEAQHRWLRPAEICEILSNFHKFHISPEPPNKPPSGSLYLFDRKVLRYFRKDGHNWRKKRDGKTVKEAHEKLKVGSVDVLHCYYAHGEDDENFQRRCY 117
5669********************************************************************************************** PP
CG-1 100 wlLeeelekivlvhylevk 118
wlLe++l +iv+vhyl+vk
Pbr028458.1 118 WLLEQDLMHIVFVHYLQVK 136
****************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 82.94 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 3.2E-80 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 3.1E-50 | 21 | 134 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 6.8E-5 | 481 | 571 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 3.11E-16 | 485 | 570 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 2.95E-16 | 664 | 780 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 19.12 | 673 | 790 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 7.56E-13 | 673 | 778 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 7.3E-17 | 673 | 784 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 11.514 | 718 | 751 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0039 | 718 | 748 | IPR002110 | Ankyrin repeat |
| Pfam | PF00023 | 7.5E-4 | 718 | 750 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 1.1 | 893 | 915 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.078 | 894 | 923 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 1.06E-7 | 894 | 944 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| Pfam | PF00612 | 0.0045 | 896 | 914 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0071 | 916 | 938 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.212 | 917 | 941 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.3E-4 | 919 | 938 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1078 aa Download sequence Send to blast |
MAERGSDAQG SRLDFRQLLV EAQHRWLRPA EICEILSNFH KFHISPEPPN KPPSGSLYLF 60 DRKVLRYFRK DGHNWRKKRD GKTVKEAHEK LKVGSVDVLH CYYAHGEDDE NFQRRCYWLL 120 EQDLMHIVFV HYLQVKGHRA NAGGIIENDE VAPDLKKGSP WTSSPSNSNG RTPSGYTDYA 180 SPSSTLTSAC EDVDSGDSHQ VSSLFDSVFE SPKMGNGPLM DNAELDPSLH PSLNNYDGQS 240 SIHGVYRPQF ENDQHRFSAD SIGVIDCQKT LGAGSWEEIL EQCTTGFHTG TSHASMSTTQ 300 IGSGEVPVEQ QTLGNSFLKE EFGSSMPFQT NWQLPLGDNS LQLPECSLDQ SMNLQRTLEA 360 NLRNTPDLVS AHLNQRNEQL VQDDLQAQLT NAESECLIIS SSEHDVSKDG NVNYAFTLRQ 420 QLLDREEGLK KVDSFSRWVS KELEEVDDLQ MQSSSGIPWS TAECGDVVDD SSLSPSISQD 480 QLFSIVDFSP KWAYTDSEIE VLVFGTFLIS QKEVTKYSWS CMFGEVEVPA QVLASGVLFC 540 FAPPHSAGQV PFYVTCSNRL ACSEVREFKY QVGSTKGLDM TDIGDGAAHE MHLHLRLESL 600 LSLRPLSPSG QLFGGVKENR NLISKIISLQ EEEDYLHAAE PTAVNYLLQN EGMEHLIKLM 660 KEKLYTWLLY KAIEDGKGPS VLDAEGQGVL HLAAALGYDW AIKPTVTAGV SINFRDVNGW 720 TALHWAAFYG RQEHTVAVLL SLGADPGALT DPSPEVPLGR TPADLASVSS HKGISGFLAE 780 SSLTTYLKSL TMNDAKEDGA AEISGTKVVK TFSERIATPV SYGDMPDALS LKDSLTAITN 840 ATQAADRIQQ MFRMQSFERK RLTEYDSDEF GVPDERAISF ITSKSHKGPV NGNAHTAAVQ 900 IQKKFRGWKK RKEFLIIRQR IVKIQAHVRG HQVRKQYKAI TWSVGILEKV ILRWRRKGTG 960 LRGFRPDAVA KIPDPQPVAS SEDDYDFLKK GRKQTEERLQ KALTRVKSMV QYPEGRAQYR 1020 RLLNVVEGFR ETKVNDMAVE VSEEKAEGSD EKAEGGDDLI DIDKLLDDDT FMSIAFD* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
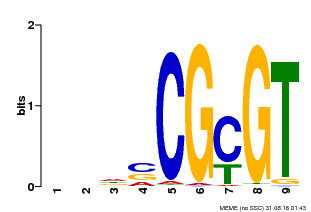
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pbr028458.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009337877.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 2 isoform X2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A498J055 | 0.0 | A0A498J055_MALDO; Uncharacterized protein | ||||
| STRING | XP_009337876.1 | 0.0 | (Pyrus x bretschneideri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7406 | 26 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



