 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pbr022669.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Pyrus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 493aa MW: 53505.1 Da PI: 5.2993 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 46.1 | 1.1e-14 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+grWT+eEde l+++++ G g+W+ ++ g+ R++k+c++rw +yl
Pbr022669.1 14 KGRWTAEEDEMLLNYIQANGEGSWRCLPKNAGLLRCGKSCRLRWINYL 61
79******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 47.3 | 4.7e-15 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg+ + +E+++++++++ lG++ W++Ia ++ gRt++++k++w+++l
Pbr022669.1 67 RGNISSQEEDIIIKLHASLGNR-WSLIAGQLP-GRTDNEIKNYWNSHL 112
78999*****************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-20 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 14.954 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.33E-27 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.8E-10 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 8.2E-13 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.76E-8 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 24.205 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.8E-24 | 65 | 116 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.3E-14 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.2E-13 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 8.33E-10 | 71 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009813 | Biological Process | flavonoid biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 493 aa Download sequence Send to blast |
MGRAPCCEKV GLKKGRWTAE EDEMLLNYIQ ANGEGSWRCL PKNAGLLRCG KSCRLRWINY 60 LRADLKRGNI SSQEEDIIIK LHASLGNRWS LIAGQLPGRT DNEIKNYWNS HLSRKIGTFR 120 RPATTTAITT EISTSSPPAG DEVSAAMELG PPKRRGGRTS RWAMKKDKTS STTKPKGLKA 180 RLRQSHHKHD NIAAAAAADD DDRLNNAHEA IALPTKSNNK NVDTMQDGYV LPMGVPDQQQ 240 ETRGGGTAMP ATVIDHQKET DGEKLILGPH PHVHGDDDMN EYVGINGGLL GFTDYLMDDI 300 NEDEILDPNG AMALSSSEIN IHQDADHFPN AVISDHQDTE LPSSCDQLVI SPNKVMKTTT 360 TTTTTTTAAS DDHHDQELES SSCGCNLSST SSISNSNGKV VTGDLHSCGS SLSLTNTINC 420 DTEDGHTAGG FGDHDQILWD WESVLSNAGT GHDYLWDGHD RQNMLSWLRE GESATAAGDG 480 DHADAMVAWL LS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 4e-23 | 12 | 118 | 5 | 110 | B-MYB |
| Search in ModeBase | ||||||
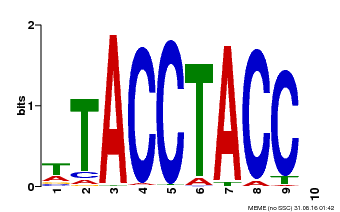
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00627 | PBM | Transfer from PK06182.1 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pbr022669.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | DQ074470 | 0.0 | DQ074470.1 Malus x domestica MYB22 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009378690.1 | 0.0 | PREDICTED: uncharacterized protein LOC103967156 | ||||
| TrEMBL | A0A498K4R8 | 0.0 | A0A498K4R8_MALDO; Uncharacterized protein | ||||
| STRING | XP_009378690.1 | 0.0 | (Pyrus x bretschneideri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF31 | 34 | 817 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47460.1 | 1e-70 | myb domain protein 12 | ||||



