 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pbr019789.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Pyrus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 426aa MW: 45790.9 Da PI: 6.2922 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 31.1 | 4.2e-10 | 272 | 320 | 3 | 55 |
HHHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 3 rahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+h+ +Er RR++i +++ L+el+P + +k Ka +L + ++Y++sLq
Pbr019789.1 272 NSHSLAERVRREKISERMRLLQELVPGC----NKITGKAVMLDEIINYVQSLQ 320
58**************************....999*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00083 | 3.43E-11 | 267 | 324 | No hit | No description |
| SuperFamily | SSF47459 | 6.67E-17 | 267 | 336 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 15.485 | 269 | 319 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 1.4E-16 | 270 | 337 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.8E-7 | 272 | 320 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 4.6E-10 | 275 | 325 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 426 aa Download sequence Send to blast |
MGGHDSEDVG FQHRNDGSLN CPSSGMKTNP LPEKVSRMAM SSVSMFKPSN GADPFFGSGW 60 DPIVSLSQSD NFGSSTGVSH GEFPNPPYPF AMENQGMSST SHLVQYPSDS SYAGMVPKLP 120 PCFGSGSFTE MVGSFGVSEC GQIANPGCPP NYNPNREGGT LTIGAQSHDD HQISEESALG 180 SSPNGKRRKR VPESNSAFSP NKNVDGELNK DMSGESSDYL KEQDEKKAKT GENAAANLRG 240 KQMGRQAKEN SQSGDPKDSY IHVRARRGQA TNSHSLAERV RREKISERMR LLQELVPGCN 300 KITGKAVMLD EIINYVQSLQ QQVEFLSMKL ATVNPELNID IERILSKDIL NSRSGGGAIF 360 GFSPGIGSPH PYPPGMFHGN LPSLLTSAPQ FPSLPQNVLD NELQGLFHMG FDSSSAIDNM 420 GQNGK* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00135 | DAP | Transfer from AT1G10120 | Download |
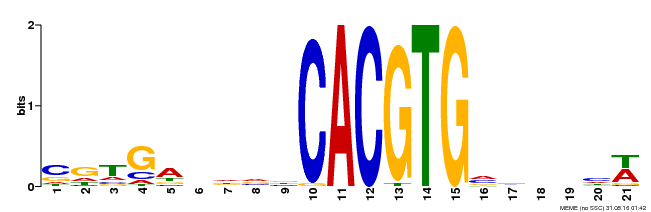
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pbr019789.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009361221.1 | 0.0 | PREDICTED: transcription factor bHLH74 isoform X2 | ||||
| TrEMBL | A0A498HQ87 | 0.0 | A0A498HQ87_MALDO; Uncharacterized protein | ||||
| STRING | XP_009361213.1 | 0.0 | (Pyrus x bretschneideri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7585 | 33 | 45 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G10120.1 | 2e-89 | bHLH family protein | ||||



