 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pbr016404.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Pyrus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 368aa MW: 42405.1 Da PI: 8.4359 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 10.7 | 0.0016 | 20 | 42 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
y C +Cg + s ks+ +Hi +H
Pbr016404.1 20 YYCDYCGICRSKKSLIASHILSH 42
89*******************96 PP
| |||||||
| 2 | zf-C2H2 | 20.1 | 1.7e-06 | 62 | 83 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
+C+ Cg +F+ + +Lk+H+++H
Pbr016404.1 62 TCEECGATFKKPAHLKQHLQSH 83
6*******************99 PP
| |||||||
| 3 | zf-C2H2 | 16.2 | 3e-05 | 89 | 113 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
y C+ dC+ s++rk++L+rH+ H
Pbr016404.1 89 YICSvdDCHSSYRRKDHLTRHLLQH 113
89*******************9887 PP
| |||||||
| 4 | zf-C2H2 | 16.4 | 2.6e-05 | 158 | 182 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C+ Cgk F+ s L++H +H
Pbr016404.1 158 HVCQeiGCGKAFRYASRLQKHEESH 182
79*******************9988 PP
| |||||||
| 5 | zf-C2H2 | 19.5 | 2.6e-06 | 249 | 273 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC C ++F++ksnL +H r H
Pbr016404.1 249 KCDykGCRRTFTTKSNLSKHVRAvH 273
688889***************9888 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50157 | 8.58 | 20 | 47 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 25 | 20 | 42 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 0.0063 | 61 | 83 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.588 | 61 | 88 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 5.1E-8 | 62 | 88 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 7.02E-10 | 62 | 113 | No hit | No description |
| PROSITE pattern | PS00028 | 0 | 63 | 83 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS50157 | 9.702 | 89 | 118 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.4E-9 | 89 | 115 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.014 | 89 | 113 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 91 | 113 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.46E-5 | 99 | 148 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-5 | 116 | 144 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.138 | 118 | 148 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.19 | 118 | 143 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 120 | 143 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 7.9E-7 | 154 | 182 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 6.52E-5 | 155 | 184 | No hit | No description |
| PROSITE profile | PS50157 | 10.907 | 158 | 187 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0086 | 158 | 182 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 160 | 182 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 8 | 190 | 215 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 192 | 215 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 15 | 218 | 239 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 8.93E-9 | 230 | 286 | No hit | No description |
| PROSITE profile | PS50157 | 13.027 | 248 | 278 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 3.9E-4 | 248 | 273 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 4.1E-7 | 249 | 273 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| Pfam | PF13894 | 0.0012 | 249 | 273 | No hit | No description |
| PROSITE pattern | PS00028 | 0 | 250 | 273 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 5.3E-5 | 274 | 302 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 5.8 | 279 | 305 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 281 | 305 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 368 aa Download sequence Send to blast |
MDKGAGEKKE GPVVIDIRRY YCDYCGICRS KKSLIASHIL SHHKEEMEMA DGGEVGEENS 60 NTCEECGATF KKPAHLKQHL QSHSLERPYI CSVDDCHSSY RRKDHLTRHL LQHQGKLFEC 120 PVENCNLKFV FQGNIKRHVN ERHTEDCPSA DCGVQKQHVC QEIGCGKAFR YASRLQKHEE 180 SHVKLDTVEA FCFEPGCMKH FTNEQCLKVH IQSCHQHTTC KICGAKQLRN NMKRHLLTHE 240 GRGSIERIKC DYKGCRRTFT TKSNLSKHVR AVHLEHKPFV CSFSGCGMRF AYKHVRDNHE 300 KTGCHVYTHG DFEEADEQFR SRPRGGRKRQ CPDIPMLVRK RVTPPNRSAQ ESAYLSWLRS 360 QEDSDEH* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1tf6_A | 3e-15 | 65 | 210 | 19 | 155 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| 1tf6_D | 3e-15 | 65 | 210 | 19 | 155 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
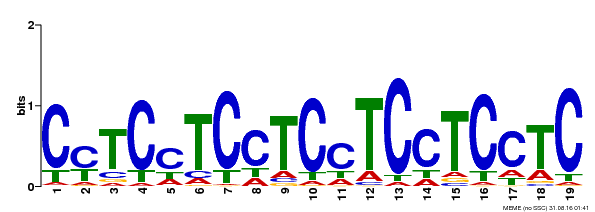
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pbr016404.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009373254.1 | 0.0 | PREDICTED: transcription factor IIIA-like isoform X1 | ||||
| Swissprot | Q84MZ4 | 1e-141 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | A0A498HWX8 | 0.0 | A0A498HWX8_MALDO; Uncharacterized protein | ||||
| STRING | XP_009373254.1 | 0.0 | (Pyrus x bretschneideri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3069 | 33 | 70 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-143 | transcription factor IIIA | ||||



