 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pbr013948.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Pyrus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 809aa MW: 87883.9 Da PI: 5.2781 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 68.4 | 9.4e-22 | 120 | 175 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
++k +++t++q++eLe++F+++++p++++r eL+++lgL+ +qVk+WFqNrR+++k
Pbr013948.1 120 KKKYHRHTPNQIQELENFFKECPHPDEKQRSELSRRLGLESKQVKFWFQNRRTQMK 175
79999************************************************999 PP
| |||||||
| 2 | START | 185 | 4.1e-58 | 327 | 546 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEE...EXCCTTEEEEEEESSS..........SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEE CS
START 2 laeeaaqelvkkalaeepgWvkss...esengdevlqkfeeskv.........dsgealrasgvvdmvlallveellddkeqWdetla....kaetle 83
la++a++elvk++++++p+W kss +++n +e+ + +ea r++ vv+ ++ lve+l+d + +W e+++ +a+t++
Pbr013948.1 327 LALAAMDELVKMVQVDSPLWIKSSdgtDILNIEEYQA------FscigmkpssLVTEATRDTCVVIVNSLALVETLMDAN-RWAEMFQclvaRASTID 417
6899********************7665555555433......333345678789*************************.***************** PP
EECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--S CS
START 84 vissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehvdlkg 174
is+g galqlm+aelq+lsp+vp R f+R+++q+ +g+w++vdvS+d +q+ p++ ++v +++lpSg+++++++n+ skvtw+eh+++++
Pbr013948.1 418 MISNGmdgtrnGALQLMYAELQMLSPMVPvRPLKFLRFCKQHAEGVWAVVDVSIDINQEGPNTNPFVDCRRLPSGCVVQDMPNNCSKVTWIEHTEYDE 515
************************************************************************************************** PP
SXXHHHHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 175 rlphwllrslvksglaegaktwvatlqrqce 205
+++h l+r +++sg+ +ga++w+atlqrq+e
Pbr013948.1 516 NIVHHLFRETLRSGMGFGAQRWLATLQRQGE 546
*****************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 9.61E-21 | 103 | 177 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.9E-22 | 107 | 177 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.945 | 117 | 177 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.0E-18 | 119 | 181 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 5.12E-20 | 120 | 177 | No hit | No description |
| Pfam | PF00046 | 3.0E-19 | 120 | 175 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 152 | 175 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 40.947 | 317 | 550 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.42E-30 | 318 | 547 | No hit | No description |
| CDD | cd08875 | 2.25E-108 | 321 | 546 | No hit | No description |
| SMART | SM00234 | 1.1E-41 | 326 | 547 | IPR002913 | START domain |
| Pfam | PF01852 | 5.3E-50 | 327 | 546 | IPR002913 | START domain |
| Gene3D | G3DSA:3.30.530.20 | 6.1E-7 | 380 | 530 | IPR023393 | START-like domain |
| SuperFamily | SSF55961 | 1.1E-21 | 568 | 802 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 809 aa Download sequence Send to blast |
MSLGGLISSS GGGNSGGGRG RVVAEIAPHA PSFMPSGTIA HPHFLSSPIP NPMQTSSSAL 60 SLSIQKMDGH SELGFAENFD PGIIGRLRDD EHESKSGSDN FEGASGDDQD GGGGDQPPRK 120 KKYHRHTPNQ IQELENFFKE CPHPDEKQRS ELSRRLGLES KQVKFWFQNR RTQMKTQLER 180 HENIILRQEN DKLIAENGVM KDAISNPVCN NCGGPAIPGQ ISFDEHQLRI ENARLKEELN 240 RICAMANKLP ISSLGITGSI LNATSGLELG VGRNGVDGMN AVSAGLPMGF DLGDGISSFS 300 PMIPPVKTSS GMMGNEVPFE RSMYIDLALA AMDELVKMVQ VDSPLWIKSS DGTDILNIEE 360 YQAFSCIGMK PSSLVTEATR DTCVVIVNSL ALVETLMDAN RWAEMFQCLV ARASTIDMIS 420 NGMDGTRNGA LQLMYAELQM LSPMVPVRPL KFLRFCKQHA EGVWAVVDVS IDINQEGPNT 480 NPFVDCRRLP SGCVVQDMPN NCSKVTWIEH TEYDENIVHH LFRETLRSGM GFGAQRWLAT 540 LQRQGESLAF LISSANSNED LSGIDVSGKK SMLKLAQRMV DNFCAGISAS SVRKWDKLCF 600 ENVSEDVRVM ARKSMNDPGE PPGIVLSVAT TVWVPVSRHR VFEFLRDEQF RKDWDILSSG 660 VPMQRIIQIG KGQEGANGVS LLRANAPNAN NSNMLILQET WTDPTGSLVV YAPVDTESIN 720 VVMRGGDSAY VALLPSGFAI LPGGPPGYEV VKSEGNGYDG SGGCLLTVAF QILVSSLPTA 780 KINMESVETV STLISCTIQR IKSALLAA* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
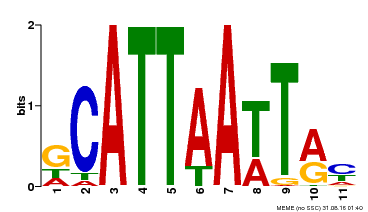
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pbr013948.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009371299.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X1 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A498JJE7 | 0.0 | A0A498JJE7_MALDO; Uncharacterized protein | ||||
| STRING | XP_009371299.1 | 0.0 | (Pyrus x bretschneideri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1192 | 34 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



