 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.J066900.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 529aa MW: 57700.7 Da PI: 5.2413 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 19.7 | 2.2e-06 | 279 | 300 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
C +Cgk F+r nL+ H+r H
Pavir.J066900.1.p 279 FCLICGKGFKRDANLRMHMRGH 300
5*******************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 7.14E-5 | 276 | 304 | No hit | No description |
| PROSITE profile | PS50157 | 11.552 | 278 | 305 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.012 | 278 | 300 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 9.9E-5 | 280 | 304 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 280 | 300 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 85 | 327 | 360 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 22 | 365 | 387 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010044 | Biological Process | response to aluminum ion | ||||
| GO:0010447 | Biological Process | response to acidic pH | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 529 aa Download sequence Send to blast |
MEGGMSSLET AMKASSSVAS GTATNADPDQ QTVRSNSVEQ FYFPRPGQSL PGIPPFFGPL 60 SSSLYLPNDN EAKVGNQFEP NPSQNTDWDP QAIVSNLTFL EQKIKQVKDV VQSMSNRENQ 120 VAGGSCELAA KQQLITADLT SIVIQLITTA GSLLPSMKNP LSSNPAVRQL SNTLGSPMGF 180 GMIANQRPSV DSKTDIPDIG KASGYEELIN SLNTTQDERD DLIKCPNPCG GEGSEPTPME 240 DLDVKESDDG EHEGENLPPG SYVVLQLEKE EILAPHTHFC LICGKGFKRD ANLRMHMRGH 300 GDEYKTAAAL AKPTKDSGSD HAPVTRYSCP FVGCKRNKEH KKFQPLETIL CVKNHYKRSH 360 CDKSYTCSRC NTKKFSVIAD LKTHEKHCGR DKWLCSCGTT FSRKDKLFGH VALFQGHTPA 420 LPMDDVKVSE ASEQQQDSEP MNEISRSMGC FPCSSSDGIS NLDMKMATPL LDPPLKMADD 480 ARGYYSPLSF DPCFGALDDF TRPGFDISED PFAFLPSGCG YVHQNGDN* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.6982 | 0.0 | callus| leaf| root| stem | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may be involved in aluminum tolerance. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00196 | ampDAP | Transfer from AT1G34370 | Download |
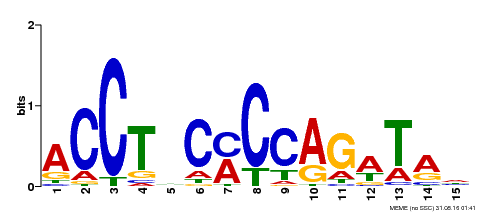
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.J066900.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KF184734 | 0.0 | KF184734.1 Saccharum hybrid cultivar R570 clone BAC 170F15 complete sequence. | |||
| GenBank | KF184941 | 0.0 | KF184941.1 Saccharum hybrid cultivar R570 clone BAC 189K21 complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025819567.1 | 0.0 | zinc finger protein STOP1 homolog | ||||
| Swissprot | Q943I6 | 0.0 | STOP1_ORYSJ; Zinc finger protein STOP1 homolog | ||||
| TrEMBL | A0A3L6RHU5 | 0.0 | A0A3L6RHU5_PANMI; Uncharacterized protein | ||||
| STRING | Pavir.J14096.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1512 | 38 | 111 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G34370.2 | 1e-153 | C2H2 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.J066900.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



