 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.7NG273800.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 370aa MW: 37618.3 Da PI: 11.1393 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 130.7 | 5e-41 | 103 | 179 | 1 | 77 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S-- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkq 77
+CqvegC+a+l+ ak+yhrrhkvC vhskap v+v g eqrfCqqCsrfh++sefD++krsCrrrLa+hnerrrk++
Pavir.7NG273800.1.p 103 RCQVEGCHAELAGAKDYHRRHKVCAVHSKAPRVVVLGAEQRFCQQCSRFHAVSEFDDAKRSCRRRLAGHNERRRKSN 179
6**************************************************************************87 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 2.2E-34 | 99 | 165 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.445 | 101 | 178 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 2.09E-38 | 102 | 181 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 1.1E-31 | 104 | 177 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0010229 | Biological Process | inflorescence development | ||||
| GO:0010321 | Biological Process | regulation of vegetative phase change | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 370 aa Download sequence Send to blast |
MHWAPAGSSP PYPQQPFVAR PGGGAASHHH HHQQQQELTC LKLGKRPCCW AGAVGTPAAQ 60 AGAALPPQVH GNGAAAGGAS GTAAEGRRKE KAGAATAAAG APRCQVEGCH AELAGAKDYH 120 RRHKVCAVHS KAPRVVVLGA EQRFCQQCSR FHAVSEFDDA KRSCRRRLAG HNERRRKSNA 180 SEAMARGAAH PHAMAPFGHG FLPPCGLPAA SPAGALSLLS SARGAGAPWL GPAPYISARS 240 SAALDELIAE NRAALLAWHF SDRLAPGRHR LAPPSSAAHH PHNGAGAWRR RPVLPRGAGQ 300 RGAHDAGPHA AAAAAATTAA PAGAPPFRPV PERAGAAARP PRTKDNGGAA GCSSDDAWAP 360 AGGGGARAL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 5e-32 | 95 | 177 | 2 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in young panicles. {ECO:0000269|PubMed:16861571}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' (By similarity). May be involved in panicle development. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00290 | DAP | Transfer from AT2G33810 | Download |
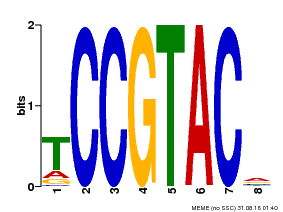
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.7NG273800.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EU964656 | 1e-110 | EU964656.1 Zea mays clone 280651 mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025824676.1 | 1e-144 | squamosa promoter-binding-like protein 7 | ||||
| Swissprot | Q7XT42 | 1e-103 | SPL7_ORYSJ; Squamosa promoter-binding-like protein 7 | ||||
| TrEMBL | A0A3L6PWC6 | 1e-157 | A0A3L6PWC6_PANMI; Squamosa promoter-binding-like protein 7 | ||||
| STRING | Pavir.Gb01121.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP10303 | 34 | 39 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G33810.1 | 2e-35 | squamosa promoter binding protein-like 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.7NG273800.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



