 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.7NG215100.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 519aa MW: 54843.6 Da PI: 7.4799 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 55.1 | 2e-17 | 131 | 220 | 1 | 86 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkm....rergf.erspkqCkekwenlnkrykkikegekkrtsessstcpyfdql 85
+Wt+ + L++a++++ e+l+rg+l++ +We+v+ ++ ++ +s +qCk+k++nl+kryk + ++ + +++ s++p+f+++
Pavir.7NG215100.1.p 131 EWTDGAISSLLDAYTDRFEQLNRGNLRGRDWEDVAGAVtdgqG-KPTgGKSVEQCKNKIDNLKKRYKVECQRLASSGAGAISHWPWFKKM 219
5*************************************65532.2223689****************999999999899*********** PP
trihelix 86 e 86
e
Pavir.7NG215100.1.p 220 E 220
8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF13837 | 5.7E-20 | 129 | 222 | No hit | No description |
| Gene3D | G3DSA:3.40.1160.10 | 8.8E-70 | 274 | 511 | IPR001048 | Aspartate/glutamate/uridylate kinase |
| SuperFamily | SSF53633 | 3.14E-51 | 276 | 511 | IPR001048 | Aspartate/glutamate/uridylate kinase |
| Hamap | MF_01220_B | 32.486 | 276 | 512 | IPR015963 | Uridylate kinase, bacteria |
| Pfam | PF00696 | 1.9E-17 | 279 | 491 | IPR001048 | Aspartate/glutamate/uridylate kinase |
| CDD | cd04254 | 8.65E-103 | 279 | 512 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006221 | Biological Process | pyrimidine nucleotide biosynthetic process | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0033862 | Molecular Function | UMP kinase activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 519 aa Download sequence Send to blast |
MASCDDDFGL LGDDAHQPAA PPPQPATAAL QPAPPPQPAQ AFCFADATVA SGAGAGAGSF 60 APAQEESNHH AERGKAAHHA KRARERPDEF SSDGGEYCSY INSGGSGGGG KKGRGGGGSS 120 GASDYRKDRE EWTDGAISSL LDAYTDRFEQ LNRGNLRGRD WEDVAGAVTD GQGKPTGGKS 180 VEQCKNKIDN LKKRYKVECQ RLASSGAGAI SHWPWFKKME QIVGNSASPA TSKALVVVED 240 EKPRQQQQHG SKRYPLSSTG PPTVVGSSRA NPLSNPRWKR VLLKIGGTAL AGAAPQNVDP 300 KIIMLIAREV QVACHHGVEV AIVVGGRNIF CGDNWAAATG TDRASTYPIG MMASVMNSVL 360 LQASLEKIGV ETRVQTALMI QEVAEPYVRR RAIRHLEKGR VVIFGGIGAG IGNPLFTTDT 420 AAALRASEIN ADVVLKGIVG DDEYGCPPRG NTNTPFEHIS FRELAARGFS RMDMTAVTCC 480 EENNIPVVIF NMLEPGNISK AICGDQIGTL VDQSGRIT* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ek5_A | 3e-60 | 270 | 511 | 1 | 239 | Uridylate kinase |
| 3ek5_B | 3e-60 | 270 | 511 | 1 | 239 | Uridylate kinase |
| 3ek5_C | 3e-60 | 270 | 511 | 1 | 239 | Uridylate kinase |
| 3ek5_D | 3e-60 | 270 | 511 | 1 | 239 | Uridylate kinase |
| 3ek5_E | 3e-60 | 270 | 511 | 1 | 239 | Uridylate kinase |
| 3ek5_F | 3e-60 | 270 | 511 | 1 | 239 | Uridylate kinase |
| 3ek6_A | 3e-60 | 270 | 511 | 1 | 239 | Uridylate kinase |
| 3ek6_B | 3e-60 | 270 | 511 | 1 | 239 | Uridylate kinase |
| 3ek6_C | 3e-60 | 270 | 511 | 1 | 239 | Uridylate kinase |
| 3ek6_D | 3e-60 | 270 | 511 | 1 | 239 | Uridylate kinase |
| 3ek6_E | 3e-60 | 270 | 511 | 1 | 239 | Uridylate kinase |
| 3ek6_F | 3e-60 | 270 | 511 | 1 | 239 | Uridylate kinase |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 108 | 116 | GGKKGRGGG |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.8413 | 0.0 | callus| leaf| root| stem | ||||
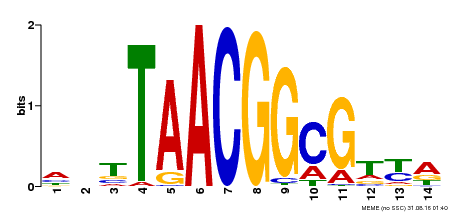
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00340 | DAP | Transfer from AT3G10030 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.7NG215100.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT067567 | 0.0 | BT067567.1 Zea mays full-length cDNA clone ZM_BFc0089D23 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025823789.1 | 0.0 | uncharacterized protein LOC112899509 isoform X1 | ||||
| TrEMBL | A0A2S3I6B5 | 0.0 | A0A2S3I6B5_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.Gb01778.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1935 | 34 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G10030.1 | 1e-159 | Trihelix family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.7NG215100.1.p |



