 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.2NG422000.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 440aa MW: 45874.4 Da PI: 7.3617 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 131.1 | 3.9e-41 | 110 | 186 | 2 | 78 |
-SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 2 CqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
C v+gC+adls++++yhrrhkvCe+hsk+pvv+v+g+e rfCqqCsrfh l efD++krsCr+rL++hn+rrrk+q+
Pavir.2NG422000.1.p 110 CAVDGCKADLSKCRDYHRRHKVCEAHSKTPVVVVAGREMRFCQQCSRFHLLAEFDDAKRSCRKRLDGHNRRRRKPQP 186
**************************************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 5.8E-36 | 102 | 171 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.129 | 107 | 184 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.24E-38 | 108 | 189 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 3.6E-31 | 110 | 183 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 440 aa Download sequence Send to blast |
MDWDLKMPVS WDLADLEHDA VPAIAAAAAA AAPPAAASSG IAAAAASARG APSRAECSVD 60 LKLGGLGEFG AADRMKEPAA PVPAGAAAPS ASPMKRPRSG AGGGAQCPSC AVDGCKADLS 120 KCRDYHRRHK VCEAHSKTPV VVVAGREMRF CQQCSRFHLL AEFDDAKRSC RKRLDGHNRR 180 RRKPQPDTMN SGSFMTSQQG LFSSAGTRFS SFPAPRPEPS WSGVIKSEDS SYYTHHQVLS 240 ARPHFAGSTP AYSKEGRRFP FLQDGDHVSF GAGAAAAAAL EVSTACQPLL KTAAAAPPHE 300 SSSSNKIFSD GLAPVLDSDC ALSLLSSPAN SSSVDVSRMV QPTEHIPMAQ PLVPSLHPPP 360 PPPPPQQQQQ FGSAPGWFAA CSQAGSGGFA CPPSVESEQL NTVLIPSSDG HEMSYHGIFH 420 VGGEGSSDGM SPSLPFSWQ* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 3e-27 | 100 | 183 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 166 | 183 | KRSCRKRLDGHNRRRRKP |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in young panicles. {ECO:0000269|PubMed:16861571}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' (By similarity). May be involved in panicle development. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00555 | DAP | Transfer from AT5G50570 | Download |
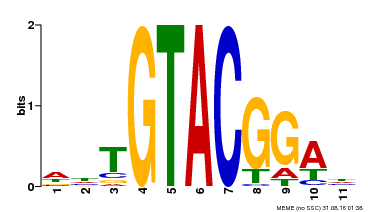
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.2NG422000.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Negatively regulated by microRNAs miR156b and miR156h. {ECO:0000305|PubMed:16861571}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT053875 | 0.0 | BT053875.1 Zea mays full-length cDNA clone ZM_BFb0379K14 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025803088.1 | 0.0 | squamosa promoter-binding-like protein 18 isoform X1 | ||||
| Swissprot | Q0J0K1 | 1e-161 | SPL18_ORYSJ; Squamosa promoter-binding-like protein 18 | ||||
| TrEMBL | A0A2S3H156 | 0.0 | A0A2S3H156_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.J30793.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1755 | 37 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G50670.1 | 4e-43 | SBP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.2NG422000.1.p |



