 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.2NG390400.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 377aa MW: 39191.5 Da PI: 9.3269 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 134.2 | 4.3e-42 | 63 | 138 | 1 | 76 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkk 76
+CqvegC +dls ak yh rhkvC++h+k+p v+v+g+eqrfCqqCsrfh+l efD++krsCrrrL +hnerrrk+
Pavir.2NG390400.1.p 63 RCQVEGCGVDLSGAKPYHCRHKVCSMHTKTPRVVVAGMEQRFCQQCSRFHQLPEFDQGKRSCRRRLIGHNERRRKP 138
6*************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.0E-34 | 59 | 125 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.394 | 61 | 138 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.7E-40 | 62 | 142 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 3.2E-32 | 64 | 137 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:2000025 | Biological Process | regulation of leaf formation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 377 aa Download sequence Send to blast |
METGGSGSNG GGSDDVRGLK FGKKIYFEQD ATAGAGGSGA AASGGRKGKG VATGPPPAAA 60 PPRCQVEGCG VDLSGAKPYH CRHKVCSMHT KTPRVVVAGM EQRFCQQCSR FHQLPEFDQG 120 KRSCRRRLIG HNERRRKPPP GPLTSRYGRL AASFQEPGRF RSFLLDFSYP RVPSSVRDAW 180 PAIQPGGDRM PGTIQWQGNQ EVHPHRSAVA GYGNHAYIGH GGSGAGPSVL PATFELPPGG 240 CVAGVATDSS CALSLLSTTQ PWDTAQSASH NRSPAMSEAS AFEGTPVAPS VMASSYPAAG 300 AWTGSRGRPA AEGARSVQHP EDALHLVHPG SVHHGNFSGE LELALQGSGP SNPPHAHHGS 360 SGGTFSHSSN AMNWSL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 9e-30 | 64 | 137 | 11 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in young panicles. {ECO:0000269|PubMed:16861571}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' (By similarity). May be involved in panicle development. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00307 | DAP | Transfer from AT2G42200 | Download |
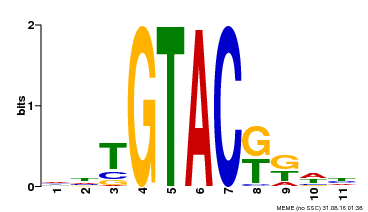
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.2NG390400.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Negatively regulated by microRNAs miR156b and miR156h. {ECO:0000305|PubMed:16861571}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025802689.1 | 0.0 | squamosa promoter-binding-like protein 17 | ||||
| Swissprot | A3C057 | 1e-157 | SPL17_ORYSJ; Squamosa promoter-binding-like protein 17 | ||||
| TrEMBL | A0A3L6TDA5 | 0.0 | A0A3L6TDA5_PANMI; Squamosa promoter-binding-like protein 17 | ||||
| STRING | Pavir.Ba01711.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2527 | 37 | 87 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G42200.1 | 2e-45 | squamosa promoter binding protein-like 9 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.2NG390400.1.p |



