 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PK15652.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Cannabis
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 371aa MW: 41007.4 Da PI: 10.1019 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 43.3 | 9e-14 | 49 | 97 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s+ykGV + +grW A+I++ ++r++lg+f ++eAa+a++ a+++++g
PK15652.1 49 SKYKGVVPQP-NGRWGAQIYEK-----HQRVWLGTFNEEDEAARAYDVAAQRFRG 97
89****9888.8*********3.....5**********99*************98 PP
| |||||||
| 2 | B3 | 104.1 | 7.5e-33 | 190 | 283 | 1 | 88 |
EEEE-..-HHHHTT-EE--HHH.HTT....---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-.SS.S CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh....ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkld.gr.s 88
f+k+ tpsdv+k++rlv+pk++ae+h ++++ +++ l +ed g++W+++++y+++s++yvltkGW++Fvk++gLk+gD+v+F+++ ++
PK15652.1 190 FEKAVTPSDVGKLNRLVIPKQHAEKHfplqSSNTSKGVLLNFEDGGGKVWRFRYSYWNSSQSYVLTKGWSRFVKEKGLKAGDIVTFQRStSNgP 283
89***************************9888899***************************************************7634444 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 1.57E-17 | 49 | 105 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 1.18E-26 | 49 | 104 | No hit | No description |
| Pfam | PF00847 | 2.8E-9 | 49 | 97 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 7.5E-21 | 50 | 105 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 2.3E-30 | 50 | 111 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 20.455 | 50 | 105 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 3.1E-41 | 184 | 295 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 4.84E-30 | 189 | 279 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.31E-30 | 189 | 282 | No hit | No description |
| Pfam | PF02362 | 2.9E-30 | 190 | 289 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 14.508 | 190 | 295 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 7.0E-25 | 190 | 295 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0048527 | Biological Process | lateral root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 371 aa Download sequence Send to blast |
XAANAAAIPP SAPKPLEKLC RLGSGASVVL DTGSSESGVE AESRKLPSSK YKGVVPQPNG 60 RWGAQIYEKH QRVWLGTFNE EDEAARAYDV AAQRFRGRDA VTNFKPLSSD TDEIEASFLS 120 SHSKAEIVDM LRKHTYNDEL EQSKRNFGAS GSPHNNRMMM MKHLLMKSSG FGSGLDDPLL 180 MKQHVREQLF EKAVTPSDVG KLNRLVIPKQ HAEKHFPLQS SNTSKGVLLN FEDGGGKVWR 240 FRYSYWNSSQ SYVLTKGWSR FVKEKGLKAG DIVTFQRSTS NGPDKQLYID WKARDNGPIV 300 GPGPVQQPKL QQHQSPPVQM LRLFGVNIFK TPGSGVVVEA GIGGGCNGKR IRDLELLELE 360 CTKKPRIIGA L |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 5e-58 | 187 | 304 | 11 | 127 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds specifically to bipartite recognition sequences composed of two unrelated motifs, 5'-CAACA-3' and 5'-CACCTG-3'. May function as negative regulator of plant growth and development. {ECO:0000269|PubMed:15040885, ECO:0000269|PubMed:9862967}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00139 | DAP | Transfer from AT1G13260 | Download |
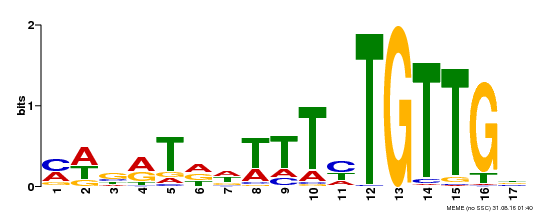
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by brassinosteroid and zeatin. {ECO:0000269|PubMed:15040885}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015895704.1 | 0.0 | AP2/ERF and B3 domain-containing transcription factor RAV1-like isoform X1 | ||||
| Refseq | XP_015895705.1 | 0.0 | AP2/ERF and B3 domain-containing transcription factor RAV1-like isoform X2 | ||||
| Swissprot | Q9ZWM9 | 1e-144 | RAV1_ARATH; AP2/ERF and B3 domain-containing transcription factor RAV1 | ||||
| TrEMBL | A0A2P5ENH5 | 0.0 | A0A2P5ENH5_TREOI; AP2/ERF transcription factor | ||||
| STRING | XP_010093403.1 | 1e-175 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2920 | 32 | 74 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G13260.1 | 1e-142 | related to ABI3/VP1 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



