 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PK04004.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Cannabis
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1079aa MW: 121872 Da PI: 5.9043 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 176.3 | 4.1e-55 | 19 | 135 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcyw 100
l+e ++rwl++ ei++iL n++ ++++ e+++rp+sgsl+L++r ++ryfrkDG++w+kkkdgktv+E+hekLKvg+v+vl+cyYah+e+n+ fqrr+yw
PK04004.1 19 LSEaQHRWLRPAEICEILRNHQTFRISPEPPSRPPSGSLFLFDRNVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSVDVLHCYYAHGEDNEYFQRRSYW 118
45559*********************************************************************************************** PP
CG-1 101 lLeeelekivlvhylevk 118
+Lee+ ++iv+vhylevk
PK04004.1 119 MLEED-KNIVFVHYLEVK 135
*****.*********985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 81.18 | 15 | 140 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.2E-76 | 18 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 3.9E-50 | 21 | 134 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 7.2E-5 | 493 | 587 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 1.7E-4 | 501 | 587 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 7.8E-14 | 501 | 587 | IPR014756 | Immunoglobulin E-set |
| Gene3D | G3DSA:1.25.40.20 | 2.8E-19 | 681 | 798 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 2.33E-20 | 685 | 796 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.76E-18 | 690 | 794 | No hit | No description |
| PROSITE profile | PS50297 | 20.367 | 690 | 806 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 8.816 | 702 | 734 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 1200 | 702 | 731 | IPR002110 | Ankyrin repeat |
| Pfam | PF12796 | 6.8E-8 | 707 | 796 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.858 | 735 | 767 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0024 | 735 | 764 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 1300 | 774 | 803 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 12 | 908 | 930 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 9.29E-8 | 908 | 959 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| PROSITE profile | PS50096 | 7.547 | 911 | 938 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0068 | 912 | 929 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.017 | 931 | 953 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.194 | 932 | 956 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 4.8E-4 | 934 | 953 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1079 aa Download sequence Send to blast |
MAERGSYGIS PRLDIQQLLS EAQHRWLRPA EICEILRNHQ TFRISPEPPS RPPSGSLFLF 60 DRNVLRYFRK DGHNWRKKKD GKTVKEAHEK LKVGSVDVLH CYYAHGEDNE YFQRRSYWML 120 EEDKNIVFVH YLEVKGNRTN VGGIRETDDS SSSLREGSPQ TSSSSTSQYK APSGSTDYTS 180 PSPNSCLTSL CEDADSGDIH QANSRLPPFA ESRQLQFGDR PLMNNLDAGF SLHHSSNHRK 240 EQSSIHVEDY IPQFKEETPN FTNPVAGSQN SLGLGTWEEI LEQCSTGYNT VPSHVSVSPS 300 QHAFVGVAHD RENVIQGKFL AREIVKEELE NSLPSDSTWQ FSSGDNVPSH LKGLVEQSSN 360 LDLSFDIGNT LFEKNTLDSN LSTPDQFSTH HGQQNERRVQ NDLQVQFTNM ESQSSMPYKF 420 DNNINTDGNV NYNFTLRQQL LDGEEGLKKV DSFSRWISKA LGDVDLQMQS SSGIPWSTVE 480 CGSADDDSSL SPSISQDQLF TIIDFSPKWA FTDSQPEVLV FGNFLKSQQE VAKYRWSCMF 540 GEVEVAAEVL AYGILCCQAP PHRDGLVPFY VTCSNRLACS ELREFNYKFG STKDIEITDI 600 YDGNTIELSL HLRLEKLLSL AVNPSSFSFK IATEKRNLIN KIISLKEEDE GYQQVDQTND 660 TGFSQYEVRD HLLTKLMKEK LYSWLLHKTI EDGKGPNILD DDGQGVLHLA AALGYDWAIK 720 PIVTAGVSIN FRDINGWTAL HWAAFYGREQ TVAYLVSLGA APGFVTDPSP EFLSGRTPAD 780 LASVNGHKGI SGFLAESSLT SHLSSLELVD SKEDGVVESS ATKAIQTVSE RTTTPRTYGE 840 MPDALSLKDS LTAVRNATQA ADRIHQMFRM QSLERRRLNE YGDDGLLDER ALSLLAGKSS 900 KAGPNDGQTY TAAVQIQKKY RGWKKRKEFL IIRQRIVKIQ AHVRGHQVRK QYRAITWSVG 960 ILEKVILRWR RKGSGLRGFR PDAVNKESVM QSLPLKEDDY DFFKEGRKQT EERLQKALTR 1020 VKSMVQYPEG RAQYRRVLNV VEGLRETKTD FTMNESEETT YDDLIEIDKL LDDDSFMYA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4hb5_A | 2e-11 | 690 | 794 | 24 | 122 | Engineered Protein |
| 4hb5_B | 2e-11 | 690 | 794 | 24 | 122 | Engineered Protein |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
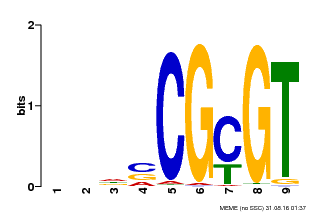
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024019885.1 | 0.0 | calmodulin-binding transcription activator 2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A2P5E8N4 | 0.0 | A0A2P5E8N4_TREOI; Notch | ||||
| STRING | XP_010094157.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7406 | 26 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



