 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PK03443.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Cannabis
|
||||||||
| Family | BBR-BPC | ||||||||
| Protein Properties | Length: 350aa MW: 39171.3 Da PI: 9.7791 | ||||||||
| Description | BBR-BPC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GAGA_bind | 384.8 | 1.4e-117 | 5 | 350 | 1 | 301 |
GAGA_bind 1 mdddgsre..rnkg.yye...........paaslkenl..glqlmssiaerdakirernlalsekkaavaerdmaflqrdkalaernkalverdnkllal 84
mdd g+re r+kg +++ +++ + +++ +q+m+++aerda+i+ernlalsekkaa+aerdmaflqrd+a+aern+a++erdn++++l
PK03443.1 5 MDDAGHREngRHKGdQFKptqgqvsgisfQNTWMMQHQptMKQIMAIMAERDAAIQERNLALSEKKAALAERDMAFLQRDAAIAERNNAMLERDNAIATL 104
99****9999****************9994433433331146********************************************************** PP
GAGA_bind 85 llvensla....salpvgvqvlsgtksidslqqlse..pqledsavelreeeklealpieeaaeeakekkkkkkrqrakkpkekkakkkkkksekskkkv 178
+++ens++ s++p+g+q+++g+k++++ q + + +++++ +++ re++++++lp++ +++a ++++ k+ +++k ++ k+ +k+ +k
PK03443.1 105 QYRENSMSngnmSSCPPGCQISRGVKHLHHPQPHVQqmQHINEASYGNREMNTNDSLPMSADTSDAVKSRRGKRIKEPKA------ISPSKRAPKPPRKI 198
******9999999*****************99944446889999**************9988777777666665444443......22333333333333 PP
GAGA_bind 179 kkesader.............................skaekksidlvlngvslDestlPvPvCsCtGalrqCYkWGnGGWqSaCCtttiSvyPLPvstk 249
kke+++ + sk+++k +dl+ln+v +Dest+P+PvCsCtG+lrqCYkWGnGGWqS+CCttt+S+yPLP +++
PK03443.1 199 KKENEELNkmtflksqewkseqsmigegdddnkqlvvSKSDWKGQDLGLNQVIYDESTMPAPVCSCTGVLRQCYKWGNGGWQSSCCTTTMSMYPLPSVPN 298
3333321123344455688899****************************************************************************** PP
GAGA_bind 250 rrgaRiagrKmSqgafkklLekLaaeGydlsnpvDLkdhWAkHGtnkfvtir 301
+r+aR++grKmS++af+klL++LaaeG+dlsnpvDLkd WAkHGtn+++ti+
PK03443.1 299 KRHARVGGRKMSGSAFNKLLSRLAAEGHDLSNPVDLKDNWAKHGTNRYITIK 350
***************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF06217 | 3.6E-102 | 5 | 350 | IPR010409 | GAGA-binding transcriptional activator |
| SMART | SM01226 | 3.8E-161 | 5 | 350 | IPR010409 | GAGA-binding transcriptional activator |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 350 aa Download sequence Send to blast |
GGFAMDDAGH RENGRHKGDQ FKPTQGQVSG ISFQNTWMMQ HQPTMKQIMA IMAERDAAIQ 60 ERNLALSEKK AALAERDMAF LQRDAAIAER NNAMLERDNA IATLQYRENS MSNGNMSSCP 120 PGCQISRGVK HLHHPQPHVQ QMQHINEASY GNREMNTNDS LPMSADTSDA VKSRRGKRIK 180 EPKAISPSKR APKPPRKIKK ENEELNKMTF LKSQEWKSEQ SMIGEGDDDN KQLVVSKSDW 240 KGQDLGLNQV IYDESTMPAP VCSCTGVLRQ CYKWGNGGWQ SSCCTTTMSM YPLPSVPNKR 300 HARVGGRKMS GSAFNKLLSR LAAEGHDLSN PVDLKDNWAK HGTNRYITIK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds to GA-rich elements (GAGA-repeats) present in regulatory sequences of genes involved in developmental processes. {ECO:0000269|PubMed:14731261}. | |||||
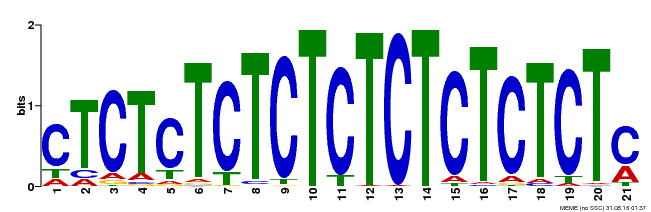
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00540 | DAP | Transfer from AT5G42520 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015884822.1 | 0.0 | protein BASIC PENTACYSTEINE6 | ||||
| Refseq | XP_015884824.1 | 0.0 | protein BASIC PENTACYSTEINE6 | ||||
| Refseq | XP_015884825.1 | 0.0 | protein BASIC PENTACYSTEINE6 | ||||
| Refseq | XP_015884826.1 | 0.0 | protein BASIC PENTACYSTEINE6 | ||||
| Refseq | XP_024930418.1 | 0.0 | protein BASIC PENTACYSTEINE6 | ||||
| Refseq | XP_024930420.1 | 0.0 | protein BASIC PENTACYSTEINE6 | ||||
| Refseq | XP_024930421.1 | 0.0 | protein BASIC PENTACYSTEINE6 | ||||
| Swissprot | Q8L999 | 1e-156 | BPC6_ARATH; Protein BASIC PENTACYSTEINE6 | ||||
| TrEMBL | A0A2P5DBV2 | 0.0 | A0A2P5DBV2_PARAD; GAGA-binding transcriptional activator | ||||
| STRING | XP_010105257.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF6905 | 34 | 49 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G42520.1 | 1e-143 | basic pentacysteine 6 | ||||



