 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PK02169.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Cannabis
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 427aa MW: 45879.6 Da PI: 10.2804 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 48.6 | 1.8e-15 | 349 | 402 | 5 | 58 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevak 58
+r+rr++kNRe+A rsR+RK+a++ eLe +v++L+++N++L+k+ ee ++ +k
PK02169.1 349 RRQRRMIKNRESAARSRARKQAYTMELEAEVAKLKEKNEELRKKQEEFVEMHKK 402
79****************************************998888777664 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 7.4E-13 | 345 | 406 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 11.254 | 347 | 396 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 2.2E-14 | 349 | 397 | No hit | No description |
| SuperFamily | SSF57959 | 1.19E-10 | 349 | 398 | No hit | No description |
| Pfam | PF00170 | 4.5E-13 | 349 | 402 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14707 | 2.79E-27 | 349 | 395 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 352 | 367 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0010255 | Biological Process | glucose mediated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 427 aa Download sequence Send to blast |
MGSNLNFKGT GNEPSANGGG GRPPANNHLV RQSSIYSLTF DEFQSTVGGI GKDFGSMNMD 60 ELLKNIWSAE EIQSMASVGP SNTGQTGVGA PLQKQGSLTL PRTLSQKTVD EVWRDLSKDG 120 SGAAGPNLPQ RQQTLGEMTL EEFLVRAGVV REDNQMAGKP PNNSGFFGDL SCSVNNHNTG 180 LGISFEQANR AATMVGNRIP DTNNQISIQA ANLPLNVNGV RSTQQHLAQQ QHQGLAQQQI 240 FPKQPTVSYA SQLPLQTGGQ LGSPGVRGGI MGITDQGLSN ALVQGGGMGM VGLGGGAVTM 300 GTGSPANQIS SDGMKSNGDT SSVSPVPFVF NGGLRGRKCS GAVEKVVERR QRRMIKNRES 360 AARSRARKQA YTMELEAEVA KLKEKNEELR KKQEEFVEMH KKQVMEMINL QREVKKRCLR 420 RTQTGPW |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 380 | 391 | KLKEKNEELRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in ABA and stress responses and acts as a positive component of glucose signal transduction. Functions as transcriptional activator in the ABA-inducible expression of rd29B. Binds specifically to the ABA-responsive element (ABRE) of the rd29B gene promoter. {ECO:0000269|PubMed:11005831, ECO:0000269|PubMed:15361142, ECO:0000269|PubMed:16284313, ECO:0000269|PubMed:16463099}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00186 | DAP | Transfer from AT1G45249 | Download |
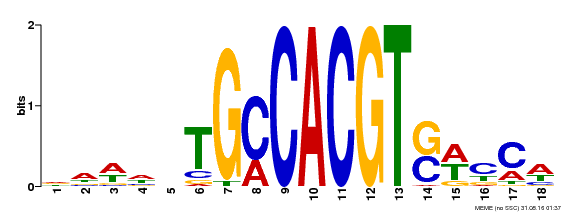
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA), cold and glucose. {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:11005831, ECO:0000269|PubMed:15361142, ECO:0000269|PubMed:16284313, ECO:0000269|PubMed:16463099}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015884344.1 | 0.0 | bZIP transcription factor TRAB1 isoform X1 | ||||
| Refseq | XP_015884345.1 | 0.0 | bZIP transcription factor TRAB1 isoform X1 | ||||
| Swissprot | Q9M7Q4 | 1e-131 | AI5L5_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 5 | ||||
| TrEMBL | A0A2P5CVH7 | 0.0 | A0A2P5CVH7_PARAD; Basic-leucine zipper transcription factor | ||||
| STRING | EOY14438 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1962 | 34 | 81 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G45249.1 | 1e-114 | abscisic acid responsive elements-binding factor 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



