 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PGSC0003DMP400026608 | ||||||||
| Common Name | LOC102605874 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1252aa MW: 139365 Da PI: 9.1029 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11.3 | 0.0011 | 1135 | 1160 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
y C C++sFs+k +L+ H ++ +
PGSC0003DMP400026608 1135 YHCDleGCSMSFSSKQELTLHKKNvC 1160
889999***************99866 PP
| |||||||
| 2 | zf-C2H2 | 11 | 0.0014 | 1159 | 1182 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirtH 23
+Cp C+k F ++ +L++H r+H
PGSC0003DMP400026608 1159 VCPveGCKKKFFSHKYLVQHRRVH 1182
69999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 10.8 | 0.0015 | 1218 | 1244 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y C+ Cg++F+ s++ rH r+ H
PGSC0003DMP400026608 1218 YACTetGCGQTFRFVSDFSRHKRKtgH 1244
889999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 1.7E-15 | 19 | 60 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.77 | 20 | 61 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 2.1E-15 | 21 | 54 | IPR003349 | JmjN domain |
| SMART | SM00558 | 4.3E-48 | 184 | 353 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 32.071 | 187 | 353 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 1.4E-24 | 198 | 367 | No hit | No description |
| Pfam | PF02373 | 2.3E-36 | 217 | 336 | IPR003347 | JmjC domain |
| SMART | SM00355 | 8.6 | 1135 | 1157 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 4.3E-5 | 1158 | 1186 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.572 | 1158 | 1187 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.012 | 1158 | 1182 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1160 | 1182 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.43E-9 | 1174 | 1216 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.0E-8 | 1187 | 1212 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.741 | 1188 | 1217 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0014 | 1188 | 1212 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1190 | 1212 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 7.38E-8 | 1206 | 1240 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 6.3E-9 | 1213 | 1240 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.157 | 1218 | 1249 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.36 | 1218 | 1244 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1220 | 1244 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1252 aa Download sequence Send to blast |
MAEASGNIEV FSWLKTLPVA PEYHPTLEEF QDPIAYIFKI EKEASKYGIC KIVPPVPEPP 60 KKTALANLNR SLSARAGSNG PTFTTRQQQI GFCPRKHRPV KKPVWQSGET YTVQQFQAKA 120 KAFEKNYLRK NSKRALTPLE VETLYWKATV DKPFSVEYAN DMPGSAFAPK KASLAAGGIG 180 EVSTLADTEW NMRGVSRSKG SLLKFMKEEI PGVTSPMVYL AMMFSWFAWH VEDHDLHSLN 240 YLHMGSGKTW YGVPRDAAVA FEEVIRVQGY AGETNPLVTF ATLGEKTTVM SPEVLLSAGI 300 PCCRLVQNAG EFVVTFPQAY HSGFSHGFNC GEASNIATPE WLRVAKDAAI RRASTNCPPM 360 VSHFQLLYDL ALSLCSRVPK NIRIEPRSSR LKDKKKSEGD MLVKELFVED LNSNNYLLHI 420 LGEGSPVVLL PQNSTGISIC SNLVAGSQSK VNSRLFPSSS SSDHEVKSKK GSAYDDLKLG 480 RKQGMEQFAG ISLEKGKYSS WHTGNRLPDS GRKDDAQSSP DTERVNLDTA RGMTYKCDTL 540 SEQGLFSCAT CGILCYTCVA IIRPTEVAAH HLMSSDYSNF NDWTGSVSGV TATGRDPNAA 600 ESDSSSGRFV KRAPALIDVP VESSDRIQKL NNGSVEGFSR TKAHKETSSL GLLALAYANS 660 SDSDEDEVEA DIPVEACESR HTDSEDEVFL RVIDPYGNHR QKRAVSQGRN CQKTDNSVQL 720 ENESYPSGES NTLLGRSSHQ PRSHQVAAKC ISNIGEIVQN NAVAPFDHAR MQFTSTSDED 780 SFRIHVFCLQ HAVQVEEQLR RIGGARISLL CHPDYPKLEA QAKQVAEELG SDHFWREISF 840 REATKDDEEM IQSALEIEEA IHGNGDWTVK LDINLFYSAN LSRSPLYSKQ MPYNFIIYNA 900 FGRNSPDNTP EKSEYTGRGS GKQRRAIVAG KWCGKVWMSS QVHPLLAERT IDEEQEQNKS 960 ISAQIKIEVK SERPRERTPT GKTVSTACKT GKKRSSTAVS RNASNAQLII ADDHDDSLLS 1020 SILQQHRKTN LRSKRIKYET PEPQKDVDKK KIFGSIIDDD PDGGPSTRLR KRIPKPSNES 1080 PAKLVKVKPA PTKQHESKKG PKVKLPSANS NAKKEPVTKG PRSNIGKRMR EEGEYHCDLE 1140 GCSMSFSSKQ ELTLHKKNVC PVEGCKKKFF SHKYLVQHRR VHMDDRPLKC PWKGCKMTFK 1200 WAWARTEHIR VHTGARPYAC TETGCGQTFR FVSDFSRHKR KTGHVSKKGR S* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 8e-72 | 13 | 441 | 8 | 413 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 8e-72 | 13 | 441 | 8 | 413 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
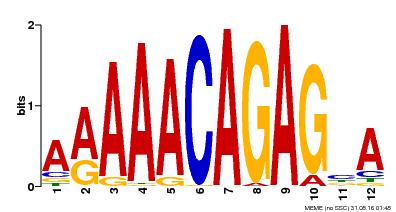
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PGSC0003DMP400026608 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG975442 | 0.0 | HG975442.1 Solanum pennellii chromosome ch03, complete genome. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006351452.1 | 0.0 | PREDICTED: lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | M1B7Y2 | 0.0 | M1B7Y2_SOLTU; Uncharacterized protein | ||||
| STRING | PGSC0003DMT400039224 | 0.0 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4988 | 23 | 31 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | PGSC0003DMP400026608 |
| Entrez Gene | 102605874 |



