 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PEQU_38228 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; Asparagales; Orchidaceae; Epidendroideae; Vandeae; Aeridinae; Phalaenopsis
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1359aa MW: 150871 Da PI: 8.9859 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 10.9 | 0.0014 | 1242 | 1265 | 1 | 22 |
EEET..TTTEEESSHHHHHHHHHH CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt 22
y+C C+ sFs+k +L H r
PEQU_38228 1242 YTCDieGCSLSFSTKQELSLHKRD 1265
99*******************985 PP
| |||||||
| 2 | zf-C2H2 | 10.7 | 0.0017 | 1325 | 1351 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y C Cg++F+ s++ rH r+ H
PEQU_38228 1325 YICRepGCGQTFRFVSDFSRHKRKtnH 1351
889999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 5.5E-17 | 19 | 60 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.118 | 20 | 61 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 4.2E-14 | 21 | 54 | IPR003349 | JmjN domain |
| PROSITE profile | PS51184 | 33.411 | 229 | 408 | IPR003347 | JmjC domain |
| SMART | SM00558 | 5.3E-46 | 239 | 408 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 2.58E-24 | 253 | 421 | No hit | No description |
| Pfam | PF02373 | 6.7E-36 | 272 | 391 | IPR003347 | JmjC domain |
| Gene3D | G3DSA:3.30.160.60 | 3.5E-4 | 1237 | 1264 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 11 | 1242 | 1264 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 8.1 | 1265 | 1289 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.221 | 1265 | 1294 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1267 | 1289 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.1E-8 | 1284 | 1317 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.094 | 1295 | 1324 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0014 | 1295 | 1319 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1297 | 1319 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 7.77E-10 | 1305 | 1355 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.5E-9 | 1318 | 1347 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.302 | 1325 | 1356 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.13 | 1325 | 1351 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1327 | 1351 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0040010 | Biological Process | positive regulation of growth rate | ||||
| GO:0045815 | Biological Process | positive regulation of gene expression, epigenetic | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0071558 | Molecular Function | histone demethylase activity (H3-K27 specific) | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1359 aa Download sequence Send to blast |
MGVEPPLVEI LPWLKSLPVA PEYHPTLSEF QDPIAYILKI EKEASAYGIC KIVPPLPAAP 60 RKTVFANLNK SFAARPQNPN SSPSTNPSSN SNSKRSPTFS TRQQQIGFCP RRPRPVQKPV 120 WQSGDCYTLS QFEAKAKLFE LSYLSKSAPA SATAASAAAT SSGSSSRKKK KIKLKKRQKK 180 SSKLFSSYLS ALEVETLYWK ACADKPFSVE YANDIPGSGF PPIGSPTGKR WREEEAASNV 240 GETAWNMRGV SRAKGSLLRF MKEEIPGVTS PMLYVAMMFS WFAWHVEDHE LHSLNYLHLG 300 AGKTWYGVPR DAALAFEDVV RIHGYCGEVN PLMTFTLLGE KTTVMSPEVL IDAGIPCCRL 360 VQNAGEFVVT FPGSYHCGFS HGFNCGEAAN IATPGWLRVA KEAAIRRAST NYPPMVSHFQ 420 LLYALALSLC SRMPICGNNE PRSSRLKDKL QGEGELMVKK TFVQSVAETN ELLSCLLEKG 480 SSCIVLPQNT SDSPLSSNSL VRSKVKLKPR LSLGLCSREE ALKASRALPC DDVMQERNTG 540 MKHLSGFGSL KGNSVAVYHR KKHSAASNNK SVTTEAASSM LELQNLKEEK DSNLEGDGLL 600 DQGLLSCVTC GILSFACVAV VEPREAASRY LTSSGCSSLF DQSFGSGEIN DVALDTKLQR 660 SDDCLGRKDG DVKDTLVEKM DHSTRYSTQD SVPIEVGSEN SKQNGLSALD LLASAYGGES 720 SDSDDERLPH EMPACTHENF ANEFLLSYRL DNDRVATKHP DLWFGKVPFA EMEQELVGAK 780 CQNAVAAENF CRTTSNDPNY LSQSADDRYL LKLDSRCYKP EHKKSLAPHS MKDSSCSSIW 840 SLGGSINSSR MEFRSSYHNS GTQDVHETNM KMQVTTPCSD DFSLCIAPCY GSDLPSENSN 900 GATQLISYHS DMKKAAVSAI QRSDKDSSRK HVFCLQHAME VEKKLCSIGG VHIMLLCHPD 960 YPKIEAEAKA LAEELGINYA WNKIDFMEAT EKDHERIQAA LKDEESIPTN SDWAVKLGIN 1020 LYYSASLSKS PLYSKQMPYN EVIYKSFSCN SPNSSMVKTK GSGKRQGRQK KIVVAGKWCG 1080 KVWMSNQVHP YLAQRKDAQD EESEEGPFAQ GGMTNPSSIN GSESNGSSAN GSSLASKKSK 1140 KRKKRPMLKS NGKRRRFNQS NDILEVIEDD DKVSQTDKGR ILRSKNRLKH KTDSEGGPSS 1200 RLRQRPMKSE DTKLVKPSFK KQNEKRAKGG QPPKSYDDEG EYTCDIEGCS LSFSTKQELS 1260 LHKRDICPVK GCGKKLFSHK YMMQHRKVHM DDRPLQCPWK GCKMTFKWAW ARTEHIRVHT 1320 GDRPYICREP GCGQTFRFVS DFSRHKRKTN HSAKKFRRS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 3e-66 | 11 | 497 | 6 | 413 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 3e-66 | 11 | 497 | 6 | 413 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1136 | 1144 | KKSKKRKKR |
| 2 | 1139 | 1143 | KKRKK |
| 3 | 1141 | 1155 | RKKRPMLKSNGKRRR |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
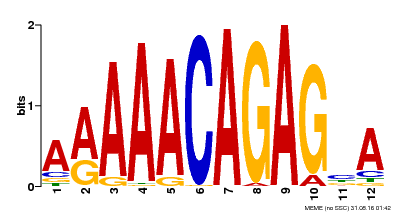
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020599580.1 | 0.0 | lysine-specific demethylase JMJ705-like | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A2I0W811 | 0.0 | A0A2I0W811_9ASPA; Lysine-specific demethylase REF6 | ||||
| STRING | XP_008797141.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5346 | 32 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



