 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PEQU_27343 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; Asparagales; Orchidaceae; Epidendroideae; Vandeae; Aeridinae; Phalaenopsis
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 373aa MW: 41950.6 Da PI: 6.8665 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 99.8 | 1.8e-31 | 248 | 303 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rF+ a++qLGG+++AtPk+i+elmkv+gLt+++vkSHLQkYRl+
PEQU_27343 248 KARRCWSPELHRRFIIALQQLGGAQVATPKQIRELMKVDGLTNDEVKSHLQKYRLH 303
68****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 13.052 | 245 | 305 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.7E-28 | 246 | 307 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 2.81E-17 | 246 | 306 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 6.5E-26 | 248 | 303 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.9E-6 | 250 | 301 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009266 | Biological Process | response to temperature stimulus | ||||
| GO:0009740 | Biological Process | gibberellic acid mediated signaling pathway | ||||
| GO:0010452 | Biological Process | histone H3-K36 methylation | ||||
| GO:0048579 | Biological Process | negative regulation of long-day photoperiodism, flowering | ||||
| GO:1903507 | Biological Process | negative regulation of nucleic acid-templated transcription | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 373 aa Download sequence Send to blast |
DGDKMGLLAP ELGLDLKLFT KRTIFSCLKD EIQAGSRAAK LEDFARRLEE ERRKIEAFKR 60 ELPLCMLLLA DVMVLKEFIP LETRSDEYKE EEKAMNLEKE SKEKMEWMSS AQLWSDNSSH 120 ESIDGKEDSQ EVFHQRTDQF VFVSSHGRKD CDALFLESNT GRAFLPFKAM AAPRKEEEKL 180 SISLPDLSLM PLPIKVPCSA PITTVAAEED NRSSGLNSKA VTIASSSPRN EHLGLQPQQQ 240 YQQNPQRKAR RCWSPELHRR FIIALQQLGG AQVATPKQIR ELMKVDGLTN DEVKSHLQKY 300 RLHTRRIPNS SATQVNRPVV MVGGLWVSQD QQNASQSGSP QGPLQLTCAA QALSMTGGDS 360 CTDDGKSESY SWK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 8e-14 | 248 | 301 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 8e-14 | 248 | 301 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 8e-14 | 248 | 301 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 8e-14 | 248 | 301 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 8e-14 | 248 | 301 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 8e-14 | 248 | 301 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 8e-14 | 248 | 301 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 8e-14 | 248 | 301 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 8e-14 | 248 | 301 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 8e-14 | 248 | 301 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 8e-14 | 248 | 301 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 8e-14 | 248 | 301 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00261 | DAP | Transfer from AT2G03500 | Download |
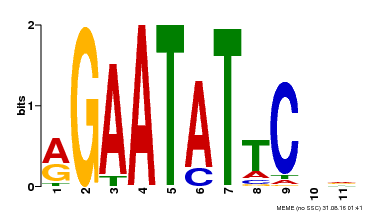
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020592704.1 | 0.0 | myb family transcription factor EFM-like isoform X1 | ||||
| TrEMBL | A0A2I0WVN2 | 0.0 | A0A2I0WVN2_9ASPA; Two-component response regulator ARR1 | ||||
| STRING | XP_008792589.1 | 1e-120 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1817 | 37 | 107 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G03500.1 | 4e-53 | G2-like family protein | ||||



