 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PEQU_08134 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; Asparagales; Orchidaceae; Epidendroideae; Vandeae; Aeridinae; Phalaenopsis
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 370aa MW: 43808 Da PI: 8.8215 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 15.1 | 6.7e-05 | 22 | 44 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
y C++Cg + s ks+L++Hi +H
PEQU_08134 22 YYCEYCGICRSKKSLLRSHILSH 44
89*******************96 PP
| |||||||
| 2 | zf-C2H2 | 17.8 | 9e-06 | 69 | 91 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C+ Cg sF + +L++H+r+H
PEQU_08134 69 HTCEECGASFWKPAHLVQHMRSH 91
79********************9 PP
| |||||||
| 3 | zf-C2H2 | 17.2 | 1.4e-05 | 97 | 121 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ Cp dC ++rk++L+rH+ +H
PEQU_08134 97 FACPveDCYFIYRRKDHLNRHLLKH 121
78*********************99 PP
| |||||||
| 4 | zf-C2H2 | 16.9 | 1.8e-05 | 126 | 150 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ Cp +C+++F+ + n++rH + H
PEQU_08134 126 FNCPveKCNRTFTFQGNMTRHVNEH 150
78*********************96 PP
| |||||||
| 5 | zf-C2H2 | 12.8 | 0.00037 | 163 | 187 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C+ Cgk F s Lk+H +H
PEQU_08134 163 HVCEepGCGKIFMFASKLKKHEDSH 187
789999***************9888 PP
| |||||||
| 6 | zf-C2H2 | 15.1 | 6.8e-05 | 254 | 278 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ C +++s ksnL++H + H
PEQU_08134 254 KCSfqGCEQTYSKKSNLNKHVKAvH 278
799999**************99877 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 10 | 22 | 44 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 9.9E-7 | 69 | 88 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0019 | 69 | 91 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 14.378 | 69 | 96 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 3.86E-10 | 69 | 121 | No hit | No description |
| PROSITE pattern | PS00028 | 0 | 71 | 91 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.5E-12 | 89 | 123 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 8.58 | 97 | 126 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.018 | 97 | 121 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 99 | 121 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.46E-6 | 108 | 157 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 6.0E-5 | 124 | 151 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.385 | 126 | 156 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.13 | 126 | 150 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 128 | 151 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.28E-5 | 159 | 189 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 5.9E-8 | 159 | 187 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 9.785 | 163 | 192 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.1 | 163 | 187 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 165 | 187 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 7.3 | 195 | 220 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 197 | 220 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 17 | 223 | 244 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 1.84E-8 | 235 | 288 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.8E-8 | 236 | 275 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.572 | 253 | 283 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0039 | 253 | 278 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 255 | 278 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS50157 | 9.058 | 284 | 313 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 1.5 | 284 | 308 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 286 | 308 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 370 aa Download sequence Send to blast |
METEAVNEDD KQRPLFRDIR RYYCEYCGIC RSKKSLLRSH ILSHHKDVFQ TVKIDEISAF 60 KAANQKVRHT CEECGASFWK PAHLVQHMRS HSVERPFACP VEDCYFIYRR KDHLNRHLLK 120 HQGKVFNCPV EKCNRTFTFQ GNMTRHVNEH HVDELPCEGK KQHVCEEPGC GKIFMFASKL 180 KKHEDSHLKV ESLEIYCGEP QCMKPFTNSE CLKAHIQSCH RYVKCEKCGT QQLRKNFKRH 240 QRLHEGVVSE ERIKCSFQGC EQTYSKKSNL NKHVKAVHEK LRPFRCRTSG CQQTFPYKHV 300 RDNHEKTHVY EQGDFLEADE RWRSRPRGGR KRERVTIEAL TRKRVVPVDC SSIMENGDEY 360 LRWLLADDNE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2i13_A | 5e-17 | 69 | 246 | 22 | 186 | Aart |
| 2i13_B | 5e-17 | 69 | 246 | 22 | 186 | Aart |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
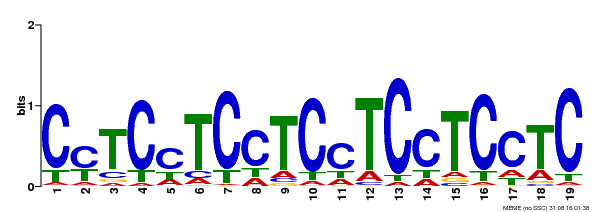
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020587074.1 | 0.0 | transcription factor IIIA isoform X1 | ||||
| Refseq | XP_020587075.1 | 0.0 | transcription factor IIIA isoform X1 | ||||
| Swissprot | Q84MZ4 | 1e-119 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | A0A2I0WK12 | 0.0 | A0A2I0WK12_9ASPA; Lysine-specific demethylase REF6 | ||||
| STRING | XP_008793643.1 | 1e-177 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2382 | 38 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-112 | transcription factor IIIA | ||||



