 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s705821g005 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | B3 | ||||||||
| Protein Properties | Length: 724aa MW: 84128.3 Da PI: 8.2817 | ||||||||
| Description | B3 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 48.7 | 1.4e-15 | 20 | 109 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT..---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh..ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsef 90
ffkvl + +++ l++p f+ ++ + ++k tl ++ g+sW+v++ ++ + + +++GW eF++a++Lk g f+vF+++g +f
PDK_30s705821g005 20 FFKVL--MHGFEEE-LRIPPAFI-KYiiD---LRGKRATLISPLGKSWSVRI--HGHNRNMCFREGWPEFARAHDLKMGFFMVFSYEGGGTF 102
78888..5566665.********.56253...35678***************..99999999***********************9885555 PP
..EEEEE-S CS
B3 91 elvvkvfrk 99
++vf++
PDK_30s705821g005 103 --SFRVFDT 109
..9999886 PP
| |||||||
| 2 | B3 | 67.7 | 1.7e-21 | 181 | 272 | 5 | 99 |
-..-HHHHTT-EE--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EEEE CS
B3 5 ltpsdvlksgrlvlpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvvkv 96
++ +l+++ + +p +f e++ + + +++++led +grsW+v+++ +++ +++ l++GW+eF+ + gL+egD ++F+l + ++ +++v++
PDK_30s705821g005 181 IIKDQNLSKNYMNIPASFRESNHIAT--NQEVILEDVEGRSWRVRIC-NRGTKGVHLAQGWQEFCADTGLEEGDKCIFELASMKDNKMLVRI 269
56778899999********9998774..459***************5.44555577999********************9988********* PP
E-S CS
B3 97 frk 99
f++
PDK_30s705821g005 270 FKR 272
986 PP
| |||||||
| 3 | B3 | 63.5 | 3.2e-20 | 348 | 437 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE.. CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefel 92
f+kv+ d s+r+++pkkf+++++ +e s+ ++l+ +sg+ W v+l ++ + +++ +GWk+Fv+++ +ke D +vFk++g+ +f
PDK_30s705821g005 348 FLKVM---DGDFSHRMTIPKKFVKNFK--EELSQNIQLKGPSGNLWHVRL--SNDAEDMIFDDGWKQFVEDHYIKEDDHLVFKYNGNLCF-- 430
56666...7788999********7776..56999****************..*********************************99999.. PP
EEEEE-S CS
B3 93 vvkvfrk 99
v +f++
PDK_30s705821g005 431 SVLIFDQ 437
8988886 PP
| |||||||
| 4 | B3 | 41.7 | 2.1e-13 | 634 | 712 | 16 | 99 |
EE--HHH.HTT---..--SEEEEEETTS.-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EEEEE-S CS
B3 16 lvlpkkfaeehggkkeesktltledesg.rsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvvkvfrk 99
++p+ fa+eh ++e++++ l+ ++g + W++ + + + g l+k Wk+F+ +n+L+egD++vF+l+g e + v++fr+
PDK_30s705821g005 634 ETIPNCFAAEHL--PRENQKIALRLPHGkKIWSMSYLH--YVGYSGLGKQWKQFALDNNLEEGDVCVFELTG--ELVMDVHIFRA 712
589********5..345669*******9788*****55..444445***********************987..66668888875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 2.0E-19 | 16 | 110 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 2.55E-20 | 16 | 110 | IPR015300 | DNA-binding pseudobarrel domain |
| PROSITE profile | PS50863 | 13.169 | 17 | 110 | IPR003340 | B3 DNA binding domain |
| CDD | cd10017 | 2.25E-19 | 18 | 108 | No hit | No description |
| Pfam | PF02362 | 5.3E-13 | 20 | 109 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 2.8E-15 | 20 | 110 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 6.8E-24 | 172 | 271 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 3.14E-20 | 175 | 273 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.60E-15 | 176 | 271 | No hit | No description |
| PROSITE profile | PS50863 | 15.509 | 177 | 273 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 2.4E-14 | 178 | 273 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 4.8E-19 | 182 | 272 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 4.5E-25 | 343 | 440 | IPR015300 | DNA-binding pseudobarrel domain |
| PROSITE profile | PS50863 | 14.833 | 345 | 438 | IPR003340 | B3 DNA binding domain |
| SuperFamily | SSF101936 | 2.55E-24 | 346 | 443 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.72E-21 | 347 | 436 | No hit | No description |
| Pfam | PF02362 | 6.5E-18 | 348 | 437 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 9.5E-17 | 348 | 438 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 4.1E-10 | 621 | 713 | IPR003340 | B3 DNA binding domain |
| SuperFamily | SSF101936 | 6.87E-19 | 632 | 711 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 4.4E-19 | 634 | 712 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 4.24E-15 | 635 | 711 | No hit | No description |
| Pfam | PF02362 | 4.7E-11 | 635 | 712 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 11.223 | 636 | 713 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005773 | Cellular Component | vacuole | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 724 aa Download sequence Send to blast |
MDPCPEMVQD RDGGAQKPEF FKVLMHGFEE ELRIPPAFIK YIIDLRGKRA TLISPLGKSW 60 SVRIHGHNRN MCFREGWPEF ARAHDLKMGF FMVFSYEGGG TFSFRVFDTN ASLKNYGPIG 120 VQTVGHEDIN QDLNSKNDSE DMIQRPEMQV SIRQMRMKDG GEMSTERKCS YCSRRKSFEA 180 IIKDQNLSKN YMNIPASFRE SNHIATNQEV ILEDVEGRSW RVRICNRGTK GVHLAQGWQE 240 FCADTGLEEG DKCIFELASM KDNKMLVRIF KRLNWNEFHK LDLVLGLSCK KSKHLREHFQ 300 KYSVLYHQTI IIKLHLRRSS GMEQRCSRCM NWEEHCYWDH FEKTRIHFLK VMDGDFSHRM 360 TIPKKFVKNF KEELSQNIQL KGPSGNLWHV RLSNDAEDMI FDDGWKQFVE DHYIKEDDHL 420 VFKYNGNLCF SVLIFDQNGC EKEACYFVKD PKDKANESCQ SGKEIMDDSL EIIHEFLPPK 480 AHSPCSSKIF CNDRKQMRRL SAPRKLPQVN DDISDNILSV KSTNPKRKIR DDNFSSDSVN 540 ENEFAICLKN VMIRSSSCLV LFLVTQNSIN YLMIQSFINY LITMSRLLAS MVCHTAWYGA 600 HGTGRKSVQN IPSSRYKLRT IRTEVYQPVP RCTETIPNCF AAEHLPRENQ KIALRLPHGK 660 KIWSMSYLHY VGYSGLGKQW KQFALDNNLE EGDVCVFELT GELVMDVHIF RAVNDTTLPS 720 RVSN |
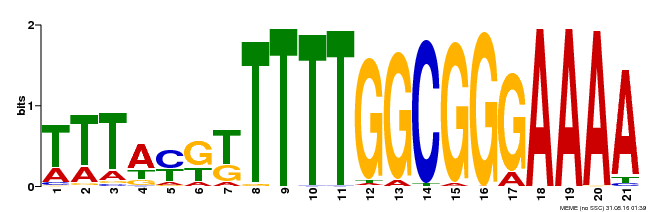
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00468 | DAP | Transfer from AT4G33280 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008786201.1 | 0.0 | putative B3 domain-containing protein Os03g0621600 | ||||
| TrEMBL | A0A2H3XN02 | 0.0 | A0A2H3XN02_PHODC; putative B3 domain-containing protein Os03g0621600 | ||||
| STRING | XP_008786201.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP562 | 34 | 179 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G33280.1 | 1e-29 | B3 family protein | ||||



