 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s660861g001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 397aa MW: 45426.3 Da PI: 5.457 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 115 | 4.8e-36 | 16 | 106 | 2 | 101 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXXXXXXXXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksFkkgkkel 93
Fl+k+y++++d+++++++sws ++ sf+v+++ efa+++LpkyFkh+nf+SFvRQLn+YgF+k++ ++ weF+++ F +g+++l
PDK_30s660861g001 16 FLTKTYDMVDDPSTNSIVSWSPTNASFIVWKPPEFARDLLPKYFKHNNFSSFVRQLNTYGFRKIDPDQ---------WEFANEDFIRGQRHL 98
9********************999*****************************************999.........*************** PP
XXXXXXXX CS
HSF_DNA-bind 94 lekikrkk 101
l++i+r+k
PDK_30s660861g001 99 LKNIHRRK 106
******98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46785 | 6.08E-36 | 11 | 105 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| Gene3D | G3DSA:1.10.10.10 | 9.3E-40 | 12 | 105 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM00415 | 2.9E-65 | 12 | 105 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 6.6E-21 | 16 | 39 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Pfam | PF00447 | 4.2E-32 | 16 | 105 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 6.6E-21 | 54 | 66 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 55 | 79 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 6.6E-21 | 67 | 79 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000302 | Biological Process | response to reactive oxygen species | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 397 aa Download sequence Send to blast |
MEGSQGGSGS NSPPPFLTKT YDMVDDPSTN SIVSWSPTNA SFIVWKPPEF ARDLLPKYFK 60 HNNFSSFVRQ LNTYGFRKID PDQWEFANED FIRGQRHLLK NIHRRKPIHS HSLQHQGNSG 120 PLAESERREL EEEIERLNCE KGLLISELQS HAQQQHGMEQ QMQSLEERLH TMEHRQRHMM 180 AFLTEIVQKP GFFSNFVQQS EFHSKKRRLP KANYFCEDAN MEEKRIMSFQ GTSSEKSDIA 240 SMHAFDIEPF EKMESSLNSL ENLFRDIDSS PELAESTSYA ESPALPLTEI HTDIRSKASE 300 IDVNAEPAAP ECDSSRDRAA GTAASTVPTG VNDVFWEQFL TETPGSSDTQ EVQSERRDAN 360 DRRSEAKMAE RGNIWWSRKN VDHLADKMGH LTSAEKT |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5d5u_B | 1e-28 | 6 | 105 | 19 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 1e-28 | 6 | 105 | 19 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 1e-28 | 6 | 105 | 19 | 129 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds DNA of heat shock promoter elements (HSE). {ECO:0000250}. | |||||
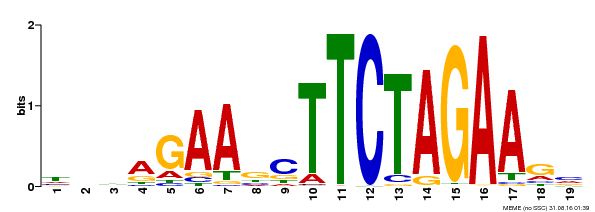
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00443 | DAP | Transfer from AT4G18880 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008776669.1 | 0.0 | heat stress transcription factor A-4b-like | ||||
| Swissprot | Q94J16 | 1e-142 | HFA4B_ORYSJ; Heat stress transcription factor A-4b | ||||
| TrEMBL | A0A2H3ZHW9 | 0.0 | A0A2H3ZHW9_PHODC; heat stress transcription factor A-4b-like | ||||
| STRING | XP_008776668.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3296 | 37 | 78 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G18880.1 | 3e-87 | heat shock transcription factor A4A | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



