 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s1042771g006 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 413aa MW: 45857 Da PI: 8.8931 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 40.2 | 6e-13 | 258 | 304 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+ L A+ YI +Lq
PDK_30s1042771g006 258 NHVEAERQRREKLNQRFYALRAVVPNiS------KMDKASLLGDAIAYITELQ 304
799***********************66......*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 2.6E-7 | 1 | 56 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| Pfam | PF14215 | 1.6E-8 | 56 | 94 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 16.85 | 254 | 303 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 6.15E-19 | 255 | 326 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 7.58E-14 | 257 | 308 | No hit | No description |
| Pfam | PF00010 | 2.1E-10 | 258 | 304 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 6.6E-19 | 258 | 317 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.1E-16 | 260 | 309 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS51671 | 9.897 | 344 | 413 | IPR002912 | ACT domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008152 | Biological Process | metabolic process | ||||
| GO:0009611 | Biological Process | response to wounding | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0016597 | Molecular Function | amino acid binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 413 aa Download sequence Send to blast |
MRKRVLQKLH VLSGGADEEI YALRLDRVTD VEMYFLSSMY FSFPRGAGGP GRALASAGFR 60 TVVLVPFDTG VLELASVKTV PESFEALQMI RAVFSPGGSN PVEMKDENGS FGERLSMAAR 120 AEERPVEIQR NGRNHHHRMA PVPAPVSSGI VNGLNWSHTQ VWAFVPHGNG VREDPRTNQF 180 STPKQQQPQP QSQLRQIDFS GGATSRASGA LVARLGVLES EHSDVEASCK EERPVAMDDR 240 RPRKRGRKPA NGREEPLNHV EAERQRREKL NQRFYALRAV VPNISKMDKA SLLGDAIAYI 300 TELQKNVKEM EAEREECPES AFMGRKRRAQ CPEIDVQAAH DEVIVRVSCP MDTHPVSKVI 360 QVFKEAQINV VDSKVTASNE AVLHTFVVKS PGSEQLMRDK LIAAISRETT SSS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 3e-29 | 252 | 314 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_B | 3e-29 | 252 | 314 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_E | 3e-29 | 252 | 314 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_F | 3e-29 | 252 | 314 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_G | 3e-29 | 252 | 314 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_I | 3e-29 | 252 | 314 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_M | 3e-29 | 252 | 314 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_N | 3e-29 | 252 | 314 | 2 | 64 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 239 | 247 | RRPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that negatively regulates jasmonate (JA) signaling (PubMed:30610166). Negatively regulates JA-dependent response to wounding, JA-induced expression of defense genes, JA-dependent responses against herbivorous insects, and JA-dependent resistance against Botrytis cinerea infection (PubMed:30610166). Plays a positive role in resistance against the bacterial pathogen Pseudomonas syringae pv tomato DC3000 (PubMed:30610166). {ECO:0000269|PubMed:30610166}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00101 | PBM | Transfer from AT2G46510 | Download |
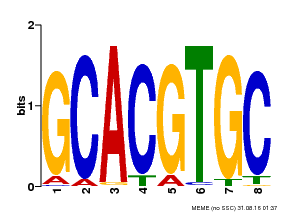
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by wounding, feeding with herbivorous insects, infection with the fungal pathogen Botrytis cinerea and infection with the bacterial pathogen Pseudomonas syringae pv tomato DC3000. {ECO:0000269|PubMed:30610166}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JX275993 | 1e-105 | JX275993.1 Cocos nucifera microsatellite WCYZ2116 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008806486.1 | 0.0 | transcription factor bHLH13-like | ||||
| Swissprot | A0A3Q7ELQ2 | 1e-136 | MTB1_SOLLC; Transcription factor MTB1 | ||||
| TrEMBL | A0A2H3Z023 | 0.0 | A0A2H3Z023_PHODC; transcription factor bHLH13-like | ||||
| STRING | XP_008806486.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5174 | 38 | 60 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46510.1 | 4e-76 | ABA-inducible BHLH-type transcription factor | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



